Cross-Over Technologies
IMAT applications constituted approximately 78 percent of all technology development applications reviewed by the NCI in FY 2005, approximately 61 percent of those reviewed in FY 2006, and 72 percent of those reviewed in FY 2007 (figure 3 below). Based on this report, 415 technology-related R21, R33, and R0l grant applications were reviewed under 5 RFAs in FY 2006, representing a growth rate of 111 percent relative to FY 2002. The explosive growth rate of such applications highlights both the success of the NCI in soliciting innovative technology development proposals and the increasing importance that innovation plays in the cancer research arena. IMAT applications constituted approximately 78 percent of all technology development applications reviewed by the NCI in FY 2005 and approximately 61 percent of those reviewed in 2006. Such data indicate the strength of the IMAT Program's contribution to the NCI's technology development portfolio. Between FY 2005 and FY 2006, the IMAT Program awarded 95 research project grants, 26 of which represented new or first-time awardees. In addition, the program has made 51 awards in FY 2007, 5 to new or first-time awardees.
Technology Applications Reviewed by the National Cancer Institute.

Technology Initiatives Applications Reviewed FY 2004 - 2008
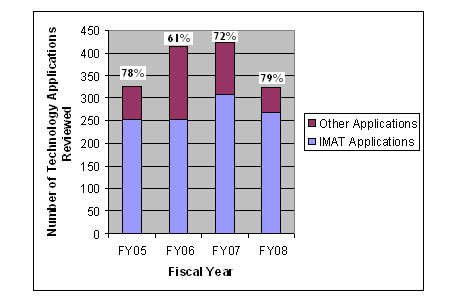
IMAT Contributions to the NCI's Technology Development Portfolio as a Function of All Technology Applications Reviewed by the National Cancer Institute
Examples of IMAT Investigators Utilizing IMAT-Developed Technologies In Selected NCI Programs
| PI Name | IMAT Award(s) | Alliance for Nanotechnology in Cancer Award | Clinical Proteomic Technologies Initiative Award | Integrative Cancer Biology Program Award |
|---|---|---|---|---|
| James Baker University of Michigan |
Photonic Crystal Fiber Probe Fluorescence Biosensing (R33 CA112141) |
DNA-linked Dendrimer Nanoparticle Systems for Cancer Diagnosis and Treatment | ||
| Xiaolian Gao University of Houston |
Parallel Synthesis of RNA Oligonucleotide Microarrays (R21 CA084708) |
Proteomic Phosphopeptide Chip Technology for Protein Profiling | ||
| Tim H-M Huang Ohio State University |
Novel Tool for Analysis of Promoter Hypermethylation (R33 CA094441) High Throughput Methylation Analysis in Cancer (R33 CA084701) |
Interrogating Epigenetic Changes in Cancer Genomes | ||
| Jan Schnitzer Sidney Kimmel Cancer Center |
Technology/Map Endothelial Targets/Human Renal Tumors (R33 CA118602) Technology to Unmask Accessible Tumor Vascular Targets (R33 CA097528) |
Nanotechnology Platform for Targeting Solid Tumors | ||
| Richard D. Smith Battelle Pacific Northwest Laboratory |
Ultra-Sensitive Approach for Monitoring Gene Production (R33 CA081654) High Throughput Fticr for Molecular Analysis of Cancer (R33 CA086340) |
A Proteomics Platform for Quantitative, Ultra-High Throughput, and Ultra-Sensitive | ||
| Paul Tempst Memorial Sloan-Kettering Cancer Center |
Peptide Profiling Techniques to Detect Thyroid Carcinoma (R21CA111942) Peptide Profiling Techniques to Detect Thyroid Carcinoma (R33CA111942) |
CPTAC team | ||
| Benjamin Cravatt Scripps Research Institute |
Microarray Platform for Profiling Cancer Proteases (R33 CA118696) | Transformative R01 (T-R01) Massively Parallel Identification of Protein Ligands |
||
| Shohei Koide University of Chicago |
High-performance affinity reagents for peptide epitopes (R21 CA132700) | Transformative R01 (T-R01) Rational Generation of Protein Capture Reagents |
||

