|
The resources developed by the Structure Group of the NCBI Computational Biology Branch (CBB) are freely available to the public and focus on four areas:
|
| |
 |
|
Macromolecular structures
The three-dimensional structures of biomolecules provide a wealth of information on their biological function and evolutionary relationships. The Molecular Modeling Database (MMDB), as part of the Entrez system, facilitates access to structure data by connecting them with associated literature, protein and nucleic acid sequences, chemicals, and more. It is possible, for example, to find 3D structures for homologs of a protein of interest by following the "Related Structure" link in an Entrez Protein sequence record (illustrated example). The 3D structures can be visualized and their sequence-structure relationships examined interactively. In addition, geometrically similar structures can be retrieved and superposed, making it possible to identify distant homologs that cannot be recognized by sequence comparison. In this way, the knowledge derived from 3D structures, which are currently available for selected protein family representatives only, may be extended to other family members.
|
|
|
|
|
 |
|
Conserved domains and protein classification
Conserved domains are functional units within a protein that have been used as building blocks in molecular evolution and recombined in various arrangements to make proteins with different functions. The Conserved Domain Database (CDD) brings together several collections of multiple sequence alignments representing conserved domains, in addition to NCBI-curated domains that use 3D-structure information explicitly to define domain boundaries and provide insights into sequence/structure/function relationships. The data are then used for putative functional annotation of protein query sequences based on matches to specific hits (illustrated example) or superfamilies, identification of proteins with similar domain architectures, and protein classification.
|
|
|
|
|
 |
|
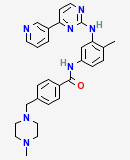
Small molecules and their biological activity
The PubChem project provides information on the biological activities of small molecules and is a component of NIH's Molecular Libraries Roadmap Initiative. PubChem includes three databases: PCSubstance, PCBioAssay, and PCCompound. The first two are archives of data submitted by the scientific community about chemical substances, including medicines, and their biological activity. The third is a derived, non-redundant database of compounds that constitute the substances in PCSubstance. The PubChem data are linked to other data types (illustrated example) in the Entrez system, making it possible, for example, to retrieve information about a compound and then "Link" to its biological activity data, retrieve 3D protein structures bound to the compound and interactively view their active sites, and find biosystems that include the compound as a component.
|
|
| |
| |
|
|
 |
|
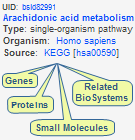
Biological Systems
A biosystem, or biological system, is a group of molecules that interact directly or indirectly, where the grouping is relevant to the characterization of living matter. The NCBI BioSystems Database provides centralized access to biological pathways from several source databases and connects the biosystem records with associated literature, molecular, and chemical data throughout the Entrez system in order to facilitate computation on biosystems data. BioSystem records list and categorize components (illustrated example), such as the genes, proteins, and small molecules involved in a biological system, along with related biosystems and citations, and allow instant retrieval of the those data sets through a wide range of Links.
|
|
| |
|
|
| |
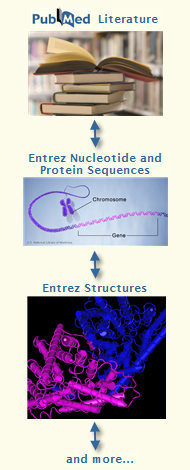
Discover associations
among previously disparate data |
| |
|
| |
|
Various data types, such as literature, nucleotide and protein sequences, and three-dimensional structures, are often submitted to public databases independently of each other by different research groups. Yet these data are related through their coverage of the same topic via different research methods. The Structure group contributes to the broader NCBI effort to identify associations among previously disparate data. See an example...
|
| |
| |
|
|



