PubChem Help
PubChem FAQ
This document provides tips and examples for searches of the three PubChem databases by text term/keyword,
as well as tips for searching PubChem Compound by chemical properties.
The help document for structure search provide tips on using chemical information for basic
and advanced structure
search options in the PubChem Structure Search.
In addition, the PubChem Deposition Gateway help
document also provides procedures and instructions for users to deposit
their structure/assay data into the PubChem system. The
PubChem Download Facility Help document
describes how to use the newly implemented PubChem download
facility.
|
PubChem Overview |

|
PubChem provides information on the biological activities of small molecules. It is a component of NIH's
Molecular Libraries
Roadmap Initiative.
PubChem includes substance information, compound structures,
and BioActivity data in three primary databases,
Pcsubstance,
Pccompound,
and PCBioAssay, respectively.
-
Pcsubstance contains more than 40 million records. You can
check the count
of substance records as of today.
- Pccompound contains more than 19 million unique structures. You can
check the count
of compound records as of today.
- PCBioAssay contains more than 1000 BioAssays. Each BioAssay contains a
various number of data points. You can
check the count
of BioAssay records as of today.
|
The Substance/Compound database, where possible, provides links to BioAssay description, literature,
references, and assay data points. The BioAssay database also includes
links back to the Substance/Compound database. PubChem is integrated
with Entrez, NCBI's primary search engine, and also provides compound neighboring, sub/superstructure, similarity structure, BioActivity data,
and other
searching features.
PubChem contains
substance and BioAssay
information from a multitude of depositors. You can check the PubChem
data source status as of today.
|
PubChem Substance Database |

|
The
PubChem substance database contains chemical structures, synonyms,
registration IDs, description, related urls, database cross-reference links to
PubMed, protein 3D structures, and biological
screening results. If the contents of a
chemical sample are known, the description includes links to
PubChem
Compound. Query Examples:
- Molecule synonym search
Which substances have "methotrexate" as a part of their molecule name?
Simply enter methotrexate in the Search
textbox on the PubChem homepage or
Entrez search page and press
the Go button. You will get all substances with
the synonym methotrexate and/or with any other
keyword methotrexate.

Or enter methotrexate[synonym] in the Search textbox and press
the Go button. Note: the
term in the brackets "[]",
such as "[synonym]", is an index field name or alias. For more information
about index searches, please see
PubChem Indexes and Index Search.
Which substances have "3'-Azido-3'-deoxythymidine" as their molecule name?
Enter "3'-Azido-3'-deoxythymidine" (including the quotes) in the
Search textbox and press the Go button.
- External ID search
Which substances have "NSC78" for DTP/NCI's external ID ?
Simply enter "NSC78" in the Search textbox and press
the Go button.
Or enter "78[objectid],dtp[sourcename]" in the Search textbox and press
the Go button.
Which substances have "aids000006" for NIAID's anti-HIV chemical
database external ID ?
Enter "aids000006" in the Search textbox and press
the Go button.
Or enter "000006[objectid],niaid[sourcename]" in the Search textbox and press
the Go button.
- Biology links search
Which substances have biological activity links?
1. Select the Limits tab or go to the
Limits page
2. In the Specify Required Links section, click the checkbox next
to BioAssay and press the Go button.
- Combined searches
Which substances contain the element Platinum and have biological activity
links?
1. Select the Limits tab or go to the
Limits page.
2. In the Specify Required Links section, click the checkbox next
to BioAssay.
3. In the Specify Required Elements section, click the checkbox to
the left of Pt.
4. Press the Go button in the Search toolbar.
| PubChem Compound Database |

|
Starting from April 6, 2005, PubChem/Compound has been updated to a new version,
and now contains unique structures only, and compound identifiers have changed.
If you are using the old version "cid"s as links, please update them.
The best way to update links to PubChem's Compound database, based on the new
cid identifiers, is simply to rerun the query or search procedure that was used
to create these links in the first place. PubChem's query and search tools
maintain all features available previously.
The
PubChem Compound Database contains validated chemical depiction information
that is provided to describe substances in
PubChem
Substance.
Structures stored within PubChem Compound are pre-clustered and
cross-referenced by identity and similarity groups. Additionally, calculated
properties and descriptors are available for searching and filtering of chemical
structures.
Users can perform a term/keyword search in a same manner as for substance
database (see above). In addition, the PubChem compound database also provides a
chemical property search.
Examples:
- Molecular weight search
Which compounds have molecular weight between 100 and 200?
Enter 100:200[mw] or 100:200[molecularweight] in the Search textbox and press
the Go button.

Note: The term in the brackets "[]",
such as "[mw]", is an index field name or alias. For more information
about index searches, please see
PubChem Indexes and Index Search.
Or simply enter 164.2[mw] in the Search textbox and press
the Go button to retrieve all compounds with 164.2 as the molecular
weight.
- XLogP search
Which compounds have XLogP between 2.3 and 2.4?
Enter 2.3:2.4[xlogp]
in the Search textbox and press
the Go button.
- Heavy atom count search (Heavy atom means all atoms except hydrogen.)
Which
compounds contain 8 heavy atoms?
Enter 8[heavyatomcount]
in the Search textbox and press
the Go button. Users can also carry out this search for the
Substance database.
The PubChem Compound Limits page provides a very useful way to rapidly
perform complex searches. All search examples showed above can be done at
the Limits page. Select the Limits tab or go to the
Limits page to begin any of the examples below.
Examples:
- Chemical property range searches
Which substances do not violate the "Lipinski Rule of 5"?
1. In the Chemical Property Search section:
a. For the Molecular Weight (MW) range, type 0 and
500 in the from and to text boxes,
respectively.
b. For the Hydrogen Bond Donor Count (HBD) range, type 0
and 5 in the from and to text boxes,
respectively.
c. For the Hydrogen Bond Acceptor Count (HBA) range, type
0 and 10 in the from and to text
boxes, respectively.
d. For the XLogP range, type -5 and 5 in the
from and to text boxes, respectively.
2. Push the Go button in the top Search bar
- Simple elemental searches of PubChem Compounds
Which substances contain Gallium?
1. In the Specify Required Elements section, select the
checkbox to the left of the Ga atomic symbol
2. Push the Go button in the top Search bar
Which substances contain Carbon, Nitrogen, Oxygen, and Fluorine?
1. In the Specify Required Elements section select the
checkboxes to the left of the C, N, O,
and F atomic symbols
2. Push the Go button in the top Search bar
|
PubChem BioAssay Database |

|
The PubChem
BioAssay Database contains BioActivity screens of chemical substances
described in PubChem
Substance. It provides searchable descriptions of each
BioAssay, including descriptions of the conditions and readouts specific
to a screening protocol.
Query Help:
- Searching for PubChem BioAssay datasets
Select
PubChem BioAssay from the pull-down menu. In
the Search textbox, enter terms you might expect to
find in the description of an assay of interest. The search will
consider terms in both the overall description of the assay and in the
description of its individual parameters and readouts.
Examples:
1. Searching for
yeast cell cycle control finds BioAssay result sets from the
NCI Yeast Anticancer Drug Screen.
2.
Searching for HIV growth inhibition finds the NCI AIDS
Antiviral Assay
- Browsing and downloading PubChem BioAssay results
The PubChem
BioAssay browser helps you to examine descriptions of each
assay's parameters and readouts. You may use it to select those
parameters and readouts most relevant to the biological activity of
interest. An example on how to work with assay data is below.
Example:
1. From the
Entrez
search page Search bar
a. Select
PubChem BioAssay from the pull-down menu.
b. Type
"NCI
AIDS Antiviral Assay" (include quote) in the
textbox.
You will see
a description of the "NCI AIDS Antiviral Assay" within
Entrez.
2. Click the hypertext link
for "AID: 179".
You will be brought to the "BioAssay Summary" page, where you will see
the detailed description of the assay. You can find more help content
about the BioAssay summary and result browser.
Query Results:
At the top of the result page, the "Tool" button  BioActivity Analysis shows the activity analysis for the BioAssays and tested compounds/substances. BioActivity Analysis shows the activity analysis for the BioAssays and tested compounds/substances.
Several popup "Link" menus are right next to the "Tool" button. These menus provide links to "Related BioAssays", "Compounds", "Literature", and other databases in Entrez. These popup links also show up for each BioAssay in the page if the BioAssay has the corresponding links.
| PubChem Summary and Analysis |

|
The
PubChem results are displayed in three category pages: substance, compound, and
BioAssay pages. They provide rich cross links to each PubChem database,
other NCBI databases, and depositor's databases. PubChem's default
results page is part of the Entrez summary list display system.
Substance Summary:
From the Entrez PubChem substance database, users can get
substance summary with thumbnails, corresponding compound ID, depositors
source information, etc. You can see
an example of a substance result in Entrez.

On this page, users can choose to display brief, summary, ID map,
substance neighboring information, synonyms, and other database information
from the dropdown list. On the right of the page, users can select few
pop-up windows (when available) to get related structure, BioAssay, and
literature links related to this substance. Users
can choose to either "display", or "send" the searched results
to "text" or to a "file".
Users can reach the more detailed substance information and cross links
by click the structure image or the ID link. Here is an example
of the PubChem Substance
Summary page:

This page displays the
standardized compound information for the substance. This page also provides a rich set
of choices for users to get property data, synonyms, descriptors, comments,
cross links, depositor's structure drawing, etc. Power users can
even download different data formats, such as ASN.1, XML, and SDF.
Compound Summary:
All compounds have been extracted from deposited substances. For natural products substances and those don't have structures, there will be no compound records
associated. A substance that is in form of mixture has the mixture format compound record and a/few component(s) compounds associated with.
From the Entrez PubChem compound database, users see a
compound summary with thumbnails, few compound property data, etc. Here
is
an example
of a compound result in Entrez.

The page is in the same style as substances. Clicking on thumbnails or CID hyperlink will lead
users to the
Compound Summary page. Users can find this compound's property
data, description, related substance information, neighboring structures, and
cross links.
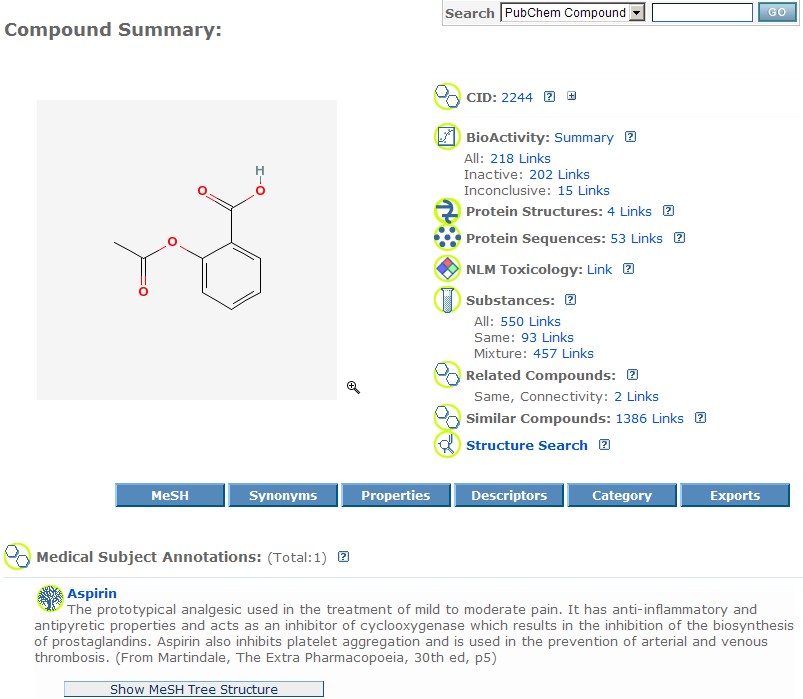
All compounds are structurally unique when compared with each other. One
compound may link to many substances.
Substance/Compound Summary Content:
Title shows chemical name and PubChem accession identifier. The toolbar
contains icons that allow users to launch: a bioactivity summary [insert
icon], when bioactivity is available; a chemical structure search [insert
icon], to search by identity, similarity, super/sub-structure, or molecular
formula; or data download in various formats, including the native PubChem
archive format ASN.1 [insert icon], XML [insert icon], or the industry
standard SDF format [insert icon].
Current Medication Info:
Content in this section is provided by the NLM DailyMed resource [http://dailymed.nlm.nih.gov/dailymed/].
DailyMed provides high quality information about marketed drugs. This
information includes FDA approved labels (package inserts). Other
information about prescription drugs may also be available.
Drug and Chemical Info:
Content in this section is provided by the NLM MeSH resource [http://www.nlm.nih.gov/mesh/].
MeSH is the U.S. National Library of Medicine's controlled vocabulary used
for indexing articles for MEDLINE/PubMed. MeSH terminology provides a
consistent way to retrieve information that may use different terminology
for the same. This section also contains pharmacological action, drug and
chemical classification, and pubmed linking information, when available.
BioActivity Results:
Content in this section is provided by the PubChem BioAssay database. A
summary of available results is provided. A launch point for bioactivity
summary analysis is provided for the current compound or the current
compound including similar compounds. To view all contributed BioAssays,
click the "more..." link.
Synonyms:
Content in this section includes all synonyms provided by depositors. The
order of the synonyms is sorted, with the most commonly used name(s) show
first. A MeSH tree icon indicates synonyms that are known to MeSH. Sorting
and display controls are available. By default, only the first ten synonyms
are shown.
Properties:
Content in this section includes computed properties of the compound record.
A list of properties are below but include various counts.
Molecular Weight -
Molecular Formula -
XLogP -
H-Bond Donor -
H-Bond Acceptor -
Rotatable Bond Count -
Tautomer Count -
Exact Mass -
MonoIsotopic Mass -
Topological Polar Surface Area
Heavy Atom Count
Formal Charge
Complexity
Isotope Atom Count
Defined Atom StereoCenter Count
Undefined Atom StereoCenter Count
Defined Bond StereoCenter Count
Undefined Bond StereoCenter Count
Covalently-Bonded Unit Count
Descriptors:
Content in this section includes computed descriptors of the compound
record. A list of descriptors is below.
IUPAC Name -
Canonical SMILES -
InChI -
Compound Info:
Content in this section is provided by the PubChem Compound database. The
PubChem Compound accession identifier (CID) is provided with the date the
CID was created and, if a mixture, the parent compound CID (when applicable)
and a link to the unique components comprising the compound record. Links to
related compounds (when applicable), with varying degrees of identity (e.g.,
being different by isotopic or stereochemical means), and 2D chemical
similarity are provided.
Substance Info:
Content in this section is provided by the PubChem Substance database. When
viewing a substance record, this section contains the PubChem Substance
accession identifier (SID) along with the dates the SID was first created
and last updated by the depositor. When the substance can be linked to a
unique compound record, the PubChem Compound accession identifier (CID) is
provided along with the date the CID was created, the parent compound CID
(when available), and a link to the unique components comprising the
compound record.
When viewing a compound record, this section contains links to all related
PubChem substance records, being either the same compound or contain the
compound as a part of a mixture. Substance Categorizations are also provided
to help you identify useful resources provided by PubChem depositors.
PubChem BioAssay Services 
A number of services are provided at PubChem to summarize the biological test results. These services include BioAssay summary for individual assay, BioActivity Summary, Data Table, and Structure-Activity Analysis for selected substance/compound/assay set. Data Table further has services for data analysis through Plots and for Selecting detailed test results. Functionality and navigation of these services are documented below.
BioAssay Summary:
BioAssay Summary may be accessed through NCBI Entrez system, where one can search the PubChem BioAssay database using a specified key word. Users can see
an example of Entrez
BioAssay search result for the term "peroxiredoxins".
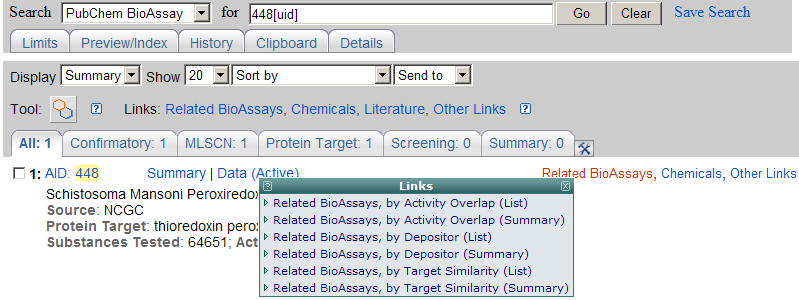
Using the "Display" pull-down menu in this page, users may
choose to view lists of summaries, brief summaries, unique identifiers,
compounds, substances, free text article links (via PMC), and PubMed
citations. On the right of the page, users can select few pop-up windows (when available) to get Related BioAssays, Compounds, Literature, etc.
Clicking on AID hyperlink will lead users to the BioAssay Summary page.

This page shows detail descriptions of a BioAssay including
citation links, experiment protocols and depositor comments. "Data Table(Active)" links to test results for compounds considered active in the particular BioAssay, while "Data Table(All)" links to the complete test results. This page also provides links to a few data analysis resources/tools that are derived at PubChem, such as "BioActivity Summary", "Related BioAssay", and etc. The bottom of the page shows detailed readouts, such as name, descriptions and data type. "Test Concentration" and "Active Concentration" attributes are flagged with * and **, respectively. The glossary of this page is listed below.
AID: PubChem's BioAssay identifier.
BioAssay Version: The BioAssay version number is composed of major version number and minor version number.
We encourage you to look at the current version result as it is the updated data from the depositors.
Name: The BioAssay name provide by the depositor.
Data Source: Depositor's source name (unique in PubChem)
Deposit Date: Date when data was first deposited.
Modify Date: Date when data was revised.
BioAssay Results: Data table for active substance or all substance.
BioActive Compounds: Active compounds/substances tested in the BioAssay. Related links for the compound/substance set.
Related BioAssays: Related BioAssays
by activity overlap, target similarity, and/or related to the same tested compound/substance set.
Protein Target: Protein target related to this
BioAssay.
Links: Extra linked information to this BioAssay.
Compounds: Compounds tested for this BioAssay, including activity information.
Substances: Substances tested for this BioAssay, including activity information.
PubMed: PubMed citations related to this BioAssay.
Protein Target: NCBI Entrez Protein links to this BioAssay if available.
Nucleotide: NCBI Entrez Nucleotide links to this BioAssay if available.
Taxonomy: NCBI Entrez Taxonomy links to this BioAssay if available.
Structure: MMDB links to this BioAssay.
Gene: Gene links to this BioAssay.
BioAssay: The BioAssays related to this one.
Description: The BioAssay's description provided by the depositor.
Protocol: The BioAssay's protocol provided by the depositor.
Comment: The BioAssay's Comment provided by the depositor.
Result Definition: The BioAssay's result definition provided by the depositors.
BioActivity Analysis
BioActivity Analysis shows the activity analysis for a set of compounds/substances and BioAssays. It has three views: Summary, Data Table, and Structure-Activity as described below.
BioActivity Analysis - Summary:This is one of the three views of "BioActivity Analysis". It displays tested compound/substance activity
summary across multiple BioAssays. The summary table contains AID, active compound/substance count, inactive
compound/substance count, discrepant count (in case of flag discrepancies option),
total tested count, outcome method, and the BioAssay name. Clicking on each count number leads to respective Data Table. Users can launch this page from PubChem Substance, Compound and BioAssay summary reports in Entrez, where users may click the display pull-down, choose
"PubChem BioActivity Summary", and see the
compound/substance activity distribution across all BioAssays. If launching from
Entrez PubChem BioAssay, users see all active compounds across each BioAssay.
Other launch points for this service are available from BioAssay Summary, Data Table and Structure-Activity Analysis services.
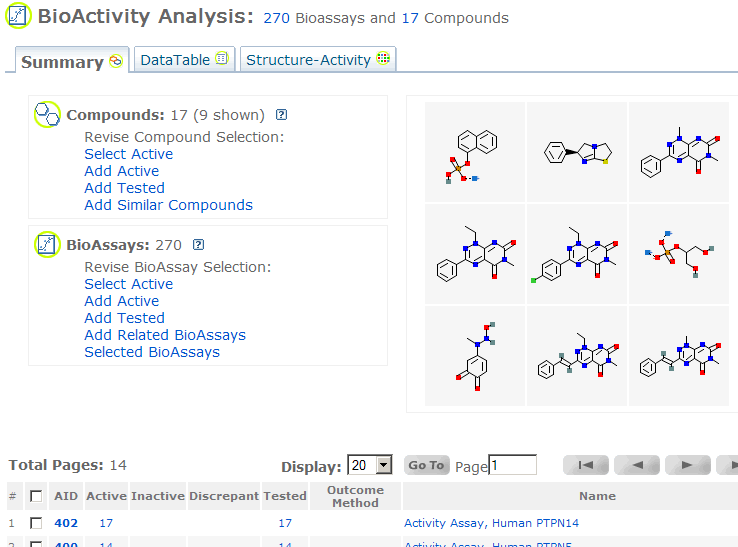
"270 BioAssays" refers to all (270 here) BioAssays in the BioActivity summary page.
"17 Compounds" refers to all compounds in the BioActivity summary page. When in substance view, users see the BioAssay and substance counts.
Data Table tab shows result data table for
selected or all (when no selection, maximum up to 50) BioAssays with the substance set in the page.
Structure-Activity tab shows Structure-Activity Analysis for
selected or all BioAssays with the substance/compound set in the page.
Revise Substance/Compound Selection allows you to reset
substance/compound based on
few options.
Select Active selects subset of active substances/compounds for
selected BioAssays.
Add Active adds all active substances/compounds for
selected BioAssays.
Add Tested adds all tested substances/compounds for
selected BioAssays.
Revise BioAssay Selection allows you to reset
BioAssays from following options.
Select Active selects subset of BioAssays in which some substances/compounds are active.
Add Active adds any BioAssay which has active substances/compounds in the
current set.
Add Tested adds any BioAssay which has tested substances/compounds in the
current set.
Add Related BioAssays adds related BioAssay by Target Similarity or by Activity Overlap.
Selected BioAssays chooses the selected and removes the rest BioAssays.
Result Display Option: Users can change the result display using the following two options.
- Group Results by: Compounds can be grouped into different levels: "Compound", "Compound, Same Connectivity", "Parent Compound", "Parent Compound, Same Connectivity". Compounds themselves are grouped from "Substances".
- Duplicate Test Option: Each compound might be tested multiple times in an assay. There are five ways to obtain a representative data: Flag Discrepancies, Exclude Duplicates, Most Recent, Most Active, and Least Active. The default is Flag Discrepancies. If a compound has multiple tested data in an assay, these data are considered "Duplicates". If the "Duplicated" data have different activity outcomes, the data are considered as "Discrepant".
Save View: is defined below.
BioActivity Analysis - Data Table:
This is one of the three views of "BioActivity Analysis". Other views "Summary" and "Structure-Activity" are available as tab options. The Data Table Page displays the searched results. There are four menus for Data Table: "Data Table, Concise", "Data Table, Complete", "Plot", and "Select".

Result Display Option: is defined above.
Save View: is defined below.
Result Exports allows you to download result set including chemical structures and readouts in the chosen format.
Data Table, Concise-- shows concise results, which contains activity outcome, score and active concentration if provided.
Data Table, Complete-- shows test results corresponding to complete or selected readout fields.
BioAssay Plot
-- This page provides an interface for plotting "Scatter Plot" and "Histogram". Users can select up to 5 rows. The "Scatter Plot" will show figures for all pairs. The "Histogram" will show figures for all rows. Users can also click on each  to get the histogram for that row. to get the histogram for that row.

Scatter Plot and Histogram: Clicking two diagonal points in the figure, you can view the data with four options: "Plot selected data", "Show selected data", "Show selected data, active only", and "Show selected data, inactive only".

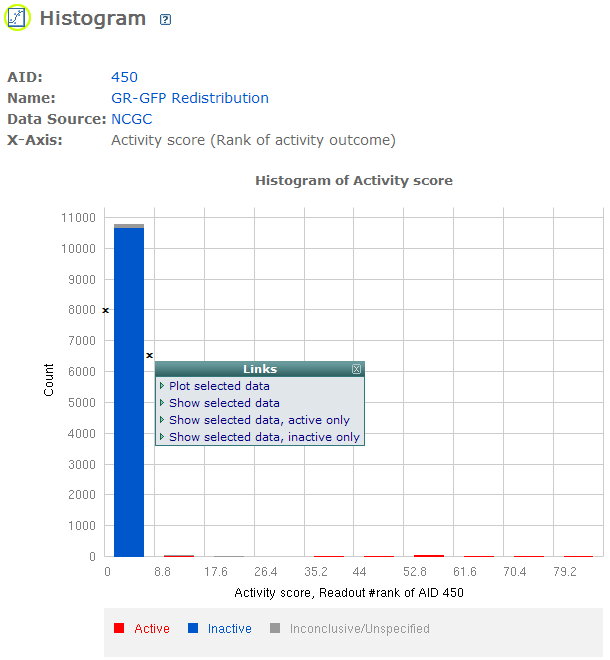
BioAssay Select
--
This page provides an interface to let you to carry out the BioAssay result search.

Navigate Buttons:
Press
the "Show" button to retrieve the BioAssay(s) data table results based on your
query criteria.
Press the "Clear" button to clear/reset the query form.
Summary Results provides a search interface for you.
You can search the activity outcome, rank score, and/or test date from the
displayed search form. Click the
 to expand the BioAssay
result search form. (Then the to expand the BioAssay
result search form. (Then the
 will be shown up. Click it will collapse the form) will be shown up. Click it will collapse the form)
Outcome Filter allows you to select tested compounds/substances based the activity outcome. The checkbox allows the outcome to be displayed in the result page. By default, it is checked.
Activity Score Filter allows you to select tested compounds/substances based the activity rank score. The checkbox allows the rank score to be displayed in the result page. By default, it is checked.
Updated Date Filter allows you to the date range for the assay. By default, all result will be returned if no input. The input format is yyyy/mm/dd. mm and dd are optional.
Other Experimental Results provides a detailed search interface for you.
Click the
 to expand the BioAssay
result search form. (Then the to expand the BioAssay
result search form. (Then the
 will be shown up. Click it will collapse the form) will be shown up. Click it will collapse the form)
All results fields are checked by default. You can unselect/select all by click the checkbox in
the header row. Selected results will be displayed in the result page.
Results with integer/float type can be searched with lower-bound value and/or upper-bound value.
String type results can be searched by either select one string term from the dropdown list or by a pattern string.
Boolean type result can be searched by select one radio button.
Pattern search: You can use pattern to perform a string search. A PATTERN is a part of a search term.
Result Filter: There are few result filters to allow you to make your result search.
Substance Filter. You can provide a SID list using list file, list text, or Entrez history to your search.
Compound Filter. You can provide a CID list using list file, list text, or Entrez history to your search.
Select Other BioAssays provide a function to allow you to add/change BioAssays.
WE DON'T ENCOURAGE YOU TO PROCESS MULTIPLE BIOASSAYS UNLESS YOU KNOW TWO OR MORE BIOASSAYS HAVE RELATION SHIP AND YOU WANT TO COMPARE THEIR RESULTS.
You can choose up to 5 BioAssays to process their data together.
BioActivity Analysis - Structure-Activity:
This is one of the three views of "BioActivity Analysis". It shows the Structure-Activity relationship in a heatmap display. The sample page is shown below. The default limit of compounds and BioAssays is set to 1000 in order to get the job done in around one minute. If more than 1000 compounds or BioAssays are input, a warning message will show up and you can change the limit to a number <= 5000. However, users need to wait for more than one minute to get the job done.

Compound/BioAssay Clusters: This is probably the most important feature in this tool. Users can cluster compounds and BioAssays differently to do the Structure-Activity analysis. Four kinds of Compounds/BioAssays relationship can be displayed: Structure/Activity, Activity/Activity, Structure/Protein Target, and Activity/Protein Target.
- Compound Cluster: can be clustered based on the structures or the activity of these compounds in the selected BioAssays. The Clustering of Compounds is shown on the left of the Heatmap. The scoring function of structure similarity is Tanimoto score, which is calculated from Structure Fingerprint. The scoring function of activity similarity is described below. The clustering algorithm for both compound and BioAssay clusters is Single Linkage.
- BioAssay Cluster: can be clustered based on the activity of the selected compounds in these BioAssays, or the sequence similarity of the Protein targets, which are the proteins interacting with the compounds in the BioAssays. The Clustering of BioAssays is shown at the top of the Heatmap. The target similarity is presented with the sequence identity.
Activity Similarity: is calculated from different data. This option is hidden by default. Users click the "+" sign near Compound Cluster to show this option.
- Calculated from: Each cell in the Heatmap corresponds to a test of one compound in one BioAssay. The result can be expressed in four kinds of data: Activity Outcome, Normalized Score - Linear, Normalized Score - Percentile, or Active Concentration if Active Concentration is available in some BioAssays. The Activity includes Active, Inactive, Unspecified/Inconclusive, or Untested. The Normalized Linear Score and Normalized Percentile Score are calculated from the raw scores of all compounds in one BioAssay. The Normalized Linear Score = ([score] - [min]) / ([max] - [min]), where [min] and [max] are the minimum and maximum score of this assay. The Normalized Percentile Score = [rank] / [N], where [rank] is the rank of one compound among all compounds in the assay, [N] is the total number of compounds in the assay. Active Concentration (AC), also called IC50, has a unit of uM. It is normalized to a value between 0 and 1 in log scale. The normalized AC = (log AC - 2) / (-3 - 2). If AC <= 0.001 uM, the normalized AC is 1. If AC > 100 uM, the normalized AC is 0.
- Scoring Functions: are different for different Data.
- Activity Outcome: The scoring function for Activity data is "Weighted Similarity" (WS). It uses both Activity Similarity of Active Compounds (ASAC) and Activity Similarity of Inactive Compounds (ASIC) data. "WS" = ("ASAC" + 0.1 * "ASIC") / (1 + 0.1). "ASAC" = [number of compounds active in both sets A and B] / ([number of compounds active in set A] + [number of compounds active in set B] - [number of compounds active in both sets A and B]). Similarly ASIC can be calculated.
- Normalized Linear score, Normalized Percentile Score, and Active Concentration: The selected scoring function for score data is "Euclidean Distance". "Euclidean Distance" = 1.0 - SUM of ([diff] * [diff]) / [N], where [N] is the total number of cells in set A (same as that in set B), [diff] = [score of a cell in set A] - [score of the corresponding cell in set B]. If both cells are untested, [diff] = 0. If one cell is tested with a score of "S" and the other cell is untested, [diff] is the higher value of "S" and 1 - "S".
Revise Selection: Users can revise both compound/substance and BioAssay. The detailed options are hidden by default. Users can click the "+" sign near Revise Selection to show the details.
Revise Compound Selection: There are three ways to modify the data set. "Select Active" selects subset of compounds/substances active in some of the selected BioAssays. "Add Active" adds additional compounds/substances active in some of the selected BioAssays. "Add Similar Compounds/Substances" adds compounds/substances similar to the current ones.
Revise BioAssay Selection: There are four ways to modify the data set. "Select Active" selects subset of BioAssays in which some of the selected compounds/substances are active. "Add Active" adds additional BioAssays in which some of the selected compounds/substances are active. "Add Related BioAssays" has two pop-up options: "by Target Similarity" and "by Activity Overlap". "Select BioAssays" has three pop-up options: "with Known Protein Targets", "Confirmatory Method", and "Summary/Confirmatory Method". "with Known Protein Targets" shows only those BioAssays with known protein targets. "Confirmatory Method" shows only those BioAssays with their Activity Outcome Method as confirmatory. "Summary/Confirmatory Method" shows only those BioAssays with their Activity Outcome Method as summary or confirmatory.
Counts: The "Counts" only appear when users launch the Heatmap for the first time. The counts include the "Input" compounds or BioAssays, the number of compounds or BioAssays with "Tested" data since some compounds or BioAssays may have no tested data in the Heatmap, the number of compounds or BioAssays "Shown" in the Heatmap.
Export:
Image: Users can export the display in one full PNG image since the display may consist of many small images.
Data Table: Users can export "BioActivity Data", "Compound Similarity Scores", and "BioAssay Similarity Scores". "BioActivity Data" shows the data corresponding to each cell in the heatmap-style display. There are three kinds of data: Activity Outcome, Score, and Active Concentration (if available). "Compound Similarity Scores" and "BioAssay Similarity Scores" show the similarity scores used to generate the dendrograms.
Clusters in GML: Users can export the clusters as a Graph Modelling Language (GML) file, which can be viewed in other softwares such as Cytoscape. The GML file format can be easily converted to other formats such as the eXtensible Graph Markup and Modeling Language (XGMML), Graph eXchange Language (GXL), and GraphML.
Result Display Option: is defined above.
Save View: is defined below.
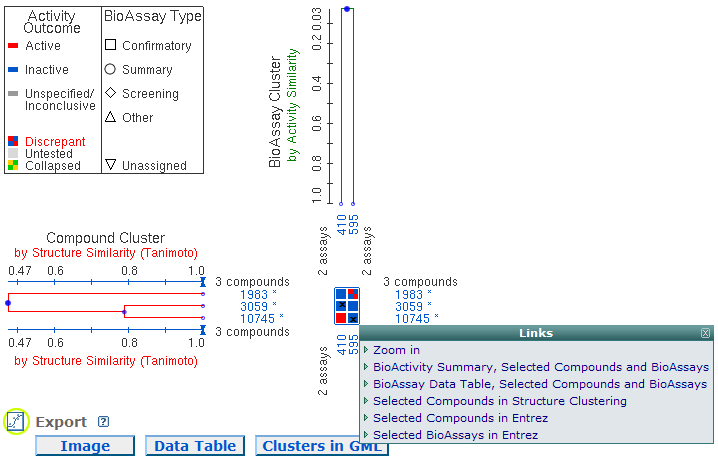
Select a region in Heatmap: One way to show a subset of the Heatmap is to click two diagonal points in the Heatmap to select compounds and BioAssays in the region as shown in the image below. A menu will pop up with six options. The first option "Zoom in" displays a new Heatmap with compounds and BioAssays in the selected region. The second option "BioActivity Summary, Selected Compounds and BioAssays" shows the selected compounds and BioAssays in PubChem BioActivity Summary page. The third option "BioAssay Data Table, Selected Compounds and BioAssays" shows the selected compounds and BioAssays in PubChem Data Table page. The fourth option "Selected Compounds in Structure Clustering" shows the selected compounds in PubChem Structure Clustering page with all structures displayed. The last two options "Selected Compounds in Entrez" and "Selected BioAssays in Entrez" show the selected compounds or BioAssays in Entrez.
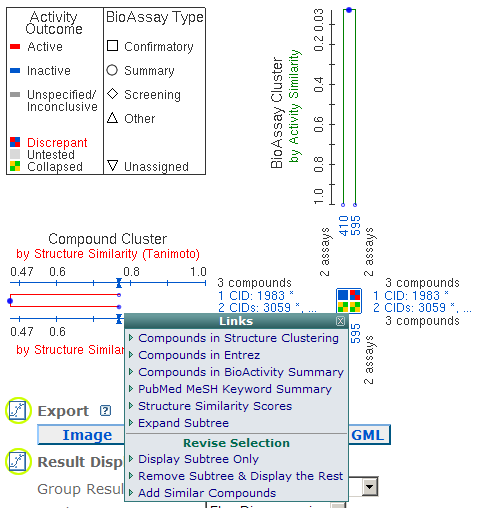
Click Blue Circles in Clusters: As shown in the following two images, if you click on a blue circle or the line above the circle in the Compound Cluster or BioAssay Cluster, a menu will pop up. The options for the Compound Cluster are "Compounds in Structure Clustering", "Compounds in Entrez", "Compounds in BioActivity Summary", "PubMed MeSH Keyword Summary" (if available), "Structure Similarity Scores" or "Activity Similarity Scores", "Expand Subtree", and three Revise Selections: "Display Subtree Only", "Remove Subtree & Display the Rest", and "Add Similar Compounds". The options for the BioAssay Cluster Tree are "BioAssays in Entrez", "BioAssays in BioActivity Summary", "Activity Similarity Scores" or "Target Sequence Identities", and four Revise Selections: "Display Subtree Only", "Remove Subtree & Display the Rest", "Add Related BioAssays, by Activity Overlap", and "Add Related BioAssays, by Target Similarity".

Collapse Compound Cluster Tree: The Compound Cluster Tree can be collapsed if users click on the ruler as shown above. The subtrees beyond the collapsed Tanimoto score will be collapsed into a node, which can be expanded. The corresponding rows in the Heatmap are collapsed as well. The color of collapsed cells is the mixture of green and yellow.
Structure Clustering:
The compounds are clustered based on the structure similarity using the Single Linkage algorithm. The structure similarity is represented using the Tanimoto score calculated from the structure fingerprints. A higher Tanimoto score means higher similarity between two structures. Both the simple view with the compound IDs (CIDs) and the view with the compound structures are provided. The limit of compounds is 5000. If more than 5000 compounds are input, a warning message will show up.
Collapse Compound Cluster: The Compound Cluster Tree can be collapsed if you click on the ruler as shown below. The subtrees beyond the collapsed Tanimoto score will be collapsed into a node, which can be expanded.
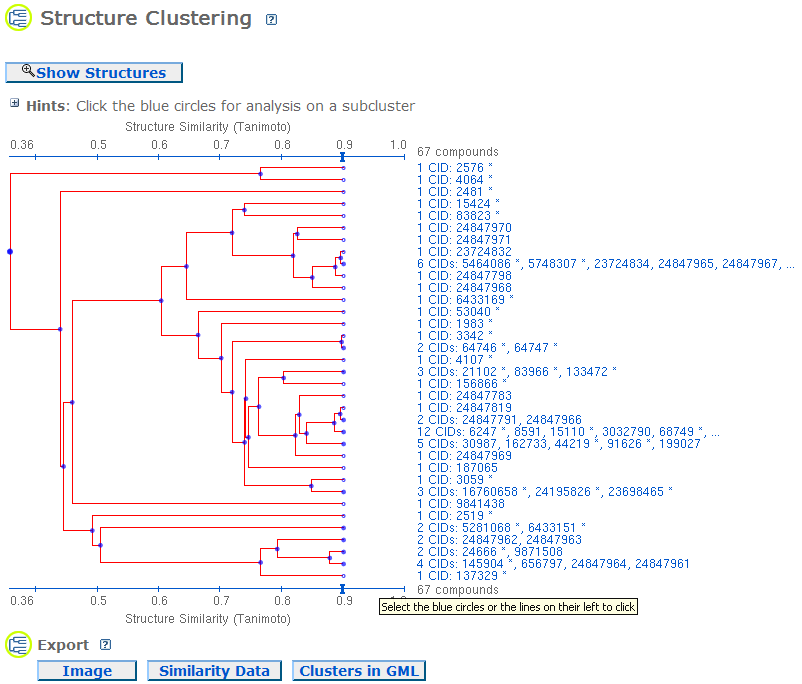

Select from Cluster: As shown in the following image, if you click on a blue circle in the Cluster Tree, a menu will pop up. The options are "Compounds in Entrez", "Compounds in BioActivity Summary", "PubMed MeSH Keyword Summary" (if available), "Structure Similarity Scores", "Expand Subtree", and two Revise Selections: "Display Subtree Only" and "Remove Subtree & Display the Rest".

Export:
Image: Users can export the display in one full PNG image since the display may consist of many small images.
Similarity Data: Users can export Structure Similarity Scores used to generate the dendrogram.
Clusters in GML: Users can export the clusters as a Graph Modelling Language (GML) file, which can be viewed in other softwares such as Cytoscape. The GML file format can be easily converted to other formats such as the eXtensible Graph Markup and Modeling Language (XGMML), Graph eXchange Language (GXL), and GraphML.
Result Display Option - Group Results by: Users can switch between "Compound" and "Substance" views. These compounds are grouped from these substances.
Save View: is defined below.
Related BioAssays by Target Similarity:
This page shows the related BioAssays based on target similarity between the queried BioAssay and all the rest BioAssays. The top 15 related BioAssays are pre-selected.

Protein Target: the protein(s) which the compounds interact with in the BioAssay.
Target Similarity: the similarity of the protein sequences for a pair of targets in two BioAssays. Both Sequence Identity and Blast E-value are shown. The related BioAssays are sorted by Sequence Identity.
Sequence Alignment: the sequences of protein targets in the original BioAssay and the Related BioAssay are aligned using Blast 2. If there are multiple targets in one BioAssay, only the target with the highest similarity is shown.
Related BioAssays by Activity Overlap:
This page shows the related BioAssays based on activity similarity between the queried BioAssay and all the rest BioAssays. The top 15 related BioAssays are pre-selected.
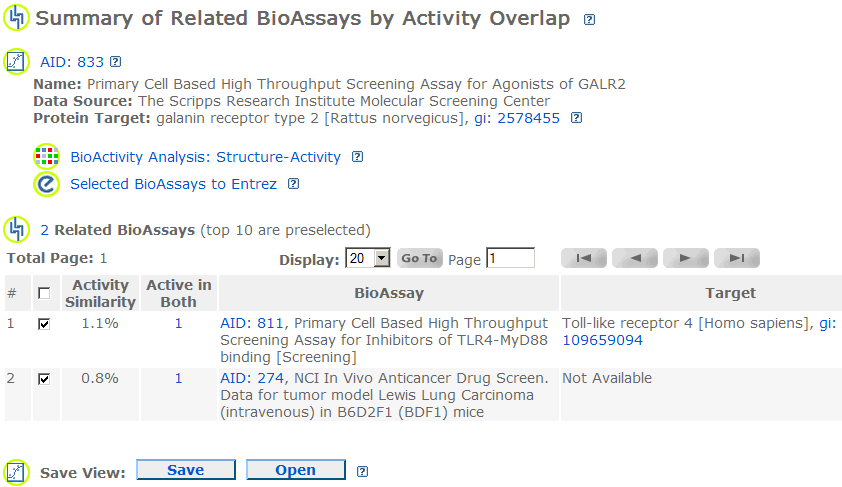
Activity Similarity: For BioAssays A and B, the activity similarity = [Active in Both A and B] / ([Active in A] + [Active in B] - [Active in Both A and B]).
Active in Both: Links to compounds active in both the queried BioAssay and the BioAssay listed in the current row.
BioAssay View File:
A BioAssay view file enables users save the state of a BioAssay display so that users may view it again at a later date or to share with colleagues. Please note that PubChem data may change over time as depositors add, update, and delete data. As such, saving a view does not absolutely guarantee that exactly the same information will be displayed at a later date. The BioAssay view file is in XML format. The specification for this file can be found at: ftp://ftp.ncbi.nih.gov/pubchem/specifications/pug.xsd.
| PubMed MeSH Keyword Summary |

|
PubMed MeSH Keyword Summary tool is intended to help users narrowing down their
PubMed searches by finding most frequent co-occurring keywords. When launched from substance/compound summary
page, the tool takes a MeSH keyword, finds all the PubMed articles associated with that keyword, and
generates a keyword summary page consisting of all the keywords found in those articles. The found keywords
are ranked by article count.
Additionally, keywords are categorized based on the parts of MeSH tree.

The displayed categories are:
Chemicals: All keywords related to "Chemicals and Drugs" except "Chemical Actions and Uses"
(keywords from D-tree besides
D-27 branch)
Pharmacological Actions of Chemicals: Pharmacological actions annotated on Chemicals and Drugs that are themselves
annotated onto PubMed articles.
Directly Annotated Pharmacological Actions: Pharmacological Actions keywords directly annotated onto PubMed articles
(keywords from D27.505 branch)
Toxicological Actions: Toxicology-related keywords (
D27.888 branch)
Biological Sciences: Keywords related to Biological Sciences (
G-branch)
Others: All other terms not included above
PubMed Articles found: the number of all articles associated with keyword(s); linked to the list of all found PubMed articles
Medical Subject keywords found: number of all keywords found; linked to a list of all MeSh keywords
Search section: the search section is for querying PubMed with a selected set of MeSH terms.
The user can select different MeSH terms by clicking on checkboxes and then adding the selected terms into the "Search"
box by clicking on "Add Selection" button. The radio buttons AND/OR/NOT are used to logically link together queries.
The default is "AND". "Clear" button clears the query box.
Search PubMed this button is for sending the query command to PubMed
Categories: All categories have keywords listed in a descending order of article count. MeSH keywords are linked to
the MeSH keyword summary page. Counts are linked to all articles associated with the keyword.
| PubChem Cross Links |

|
PubChem provides cross links to other databases when that information is available. You can find those links from either Entrez PubChem pages or individual record summary pages.
SID: Link to Entrez Pcsubstance.
Substance Version: The dropdown action displays substance information
from old version if available.
CID: Link to Entrez Pccompound.
Substances Links: (on the compound summary page)
All: Links to Entrez Pcsubstance records with all substances that contain this structure.
Same: Links to Entrez Pcsubstance records with those that contain this structure only.
Mixture: Links to Entrez Pcsubstance records where this molecule is a component of a mixture.
Component: Links to Entrez Pccompound records with component records only.
BioAssay Information: Links to the Entrez Pcassay or the BioAssay
summary page related to this compound/substance.
Related Compounds/Substances Links:
Same: The molecules in this group are exactly the same, including connectivity, isotopes and stereochemistry.
Same, Connectivity: The molecules in this group have the same regular chemical connectivity, ignoring isotopes and stereochemistry.
Same, Stereochemistry: Molecules that have the same connectivity and stereochemistry, ignoring isotopes.
Same, Isotope: Molecules that have the same connectivity and isotopes, ignoring stereochemistry.
Same, Any Tautomers: Those molecules that are tautomers.
Parent Compound Link: Link to the parent compound of the record.
A parent is conceptually the "important" part of the molecule when the molecule has more then one covalent component. Specifically, a parent component must have at least one carbon and contain at least 70% of the heavy (non-hydrogen) atoms of all the unique covalent units (ignoring stoichiometry). Note that this is a very empirical definition and is subject to change.
Same Parent Links: Links to records that share the same parent component. This is a sort of merger of the parent and related concepts: these are links to groups that share the same parent at different levels of "sameness" as described above for the Related links.
Similar Compounds Link: (on the
compound summary page) Link to Entrez
Pccompound records. All compounds shown have a similarity score [Tanimoto]
>90%.
If you want to find compounds with different scores, you can visit the PubChem
structure search page.
Similar Substances Link: (on the
substance summary page) Link to Entrez
Pcsubstance records. All substances shown have a similarity score [Tanimoto]
>90%.
If you want to find substances with different scores, you can visit the PubChem
structure search page.
Similarity links are pre-computed in PubChem using a dictionary-based
fingerprint
at 90% using the Tanimoto score equation:
Tanimoto = AB / ( A + B - AB )
Where:
Tanimoto is the Tanimoto score, a fraction between 0 and 1.
AB is the count of bits set after bit-wise & of fingerprints A and B
A is the count of bits set in fingerprint A
B is the count of bits set in fingerprint B
Each similarity link is equivalent to a chemical structure
similarity search of the PubChem Compound database yielding all chemical
structures with a Tanimoto score that is 90% or above.
In addition to the Tanimoto equation above, PubChem uses a "boost" scheme
that assigns a similarity score of: 104%
to structures with identical stereo, isotope, and connectivity.
103% to structures with identical connectivity
and either stereo or isotope.
102% to structures with identical connectivity.
101% to structures that are tautomers of the
query.
The cases of "boosted" scores greater than 101% correspond to cases
that originally would have had a score of 100% similarity. However, in the
case where tautomers get an artificial score of 101%, their natural score
could be much lower, sometimes as low as 60%, especially for small
compounds where the tautomeric system is a large part of the structure.
There are 881 substructure-keys (skeys) in each fingerprint. Each bit
in the fingerprint represents the presence (or absence) of a particular
chemical substructure (e.g., a carboxylic acid) or a particular count of
the same. These skeys are similar in nature to the well-known MDL MACCS
skeys fingerprints.
|
Structure Search: Leads you to the
PubChem structure search page by transferring this compound's isomeric SMILES
string into the search field. For more information about the structure
search, visit the structure search help page.
BioActivity: Link to the BioAssay data summary page.
Structure Links: Link to the Entrez Structure database with associated MMDB
IDs
provided by depositors.
PubMed: Links to PubMed references with this
compound/substance. The linkage is mainly based on the pmid(s) associated
with the substance/compound.
Nucleotides: Linking to the Entrez nucleotides
database with associated nucleotides GIs provided by depositors.
Protein: Links to the Entrez protein database with associated protein
GIs
provided by depositors.
Source and Source-ID: Links to the depositor's original information page by
the depositor's source name and/or source-id (external id), if available.
Medical Subject Annotations (MeSH): Linking information to records in NLM's
Medical Subject Heading (MeSH) database. Linkage is based on matching names
and synonyms supplied with the chemical structure record to those in the MeSH
record. The names and synonyms creating a link to each MeSH record are
indicated by "MeSH" links in the "Synonyms" section of PubChem
Compound and Substance Summary displays.
This section also provides PubMed information through MeSH heading and subheadings. To learn more about MeSH, visit
the MeSH site.
NLM Toxicology Link: Links to the SIS/ChemIDPlus record for this compound/substance,
which provides additional links to toxicology information sources.
More information
is available.
| PubChem Indexes and Filters in Entrez |

|
The PubChem index search is a very powerful tool within the Entrez system. Users can
simply type search term(s) followed by the bracketed index field name. Then click
the "Go" button.
Examples:
Search for DTP/NCI's record with NSC#78:
On the PubChem homepage or
Entrez search page,
enter "DTP/NCI[Sourcename], 78[objectid]" in
the search box, then click the Go button.
Search for all substances containing gold:
On the PubChem homepage or
Entrez search page
enter "Au[el]", and click the Go button.
Search for all compounds with heavy atom count between 10 and 12:
On the PubChem homepage or
Entrez
search page, choose 'Pccompound' database from the
search dropdown list,
enter "10:12[hac]", and click the Go button.
The following fields can be searched within
Entrez PubChem databases (with field
aliases in square brackets; pick one alias that's easily memorized in case
multiple aliases are available). For integer/real number fields, the range search
can be done as shown above.
PubChem Compound:All [ALL]: All of the following fields are searched. If a string query is presented without
a field alias, by default, [ALL] is searched.
Uid [UID]: The integer represents CID for each Pccompound
database. By default, an integer without a field alias is recognized as a UID. Same as [CID].
Filter [Filter]: Limits the records. A number of filters are available to restrict the search to compounds
with particular information. The specialized Filters in this database are:
- has_mesh: records with associated MeSH terms
- has_pharm: records with associated pharmacological actions
- has_parent: records that have a parent structure
- has_no_parent: records that do not have a parent
ActiveAid [AA]: Active BioAssay identifier, integer. This choice is intended for users querying for active
compounds for a particular assay. BioAssay ID (AID) should be provided.
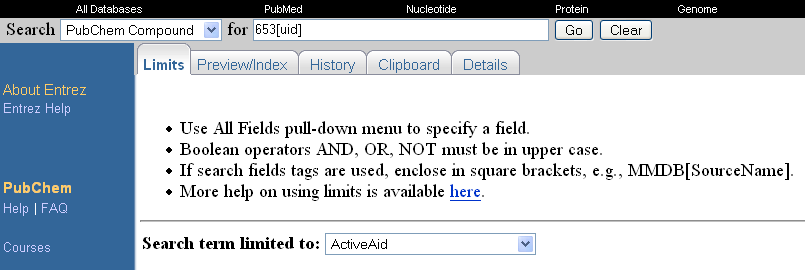
ActiveAidCount [AC, ACNT]: Using this filter users can query for compounds which are active in a certain number of assays
ActiveAidRatio [AAR]: Ratio should be between zero and 1. Ratio equals to the number of BioAssays where compounds were tested
active divided by number of BioAssays where compounds tested with any result.
ActiveConcentration: Using this filter users can query for compounds which show certain extent of active concentration(uM) in biological assays.
AssaySourceName [ASRC, ASRCNAM, ASRCNAME]: Allows filtering of by assay source name. For available data sources look
here
AtomChiralCount [ACC, ACCNT]: Total count of chiral atoms in a given
compound, integer.
AtomChiralDefCount [ACDC, ACDCNT]: Total count of defined chiral
atoms in a given compound, integer.
AtomChiralUndefCount [ACUC, ACUCNT]: Total count of undefined
chiral atoms in a given compound, integer.
BioAssayID [BAID, AID]: BioAssay identifier, integer.
BondChiralCount [BCC, BCCNT]: Total count of chiral bonds in a given compound, integer.
BondChiralDefCount [BCDC, BCDCNT]: Total count of defined chiral bonds in a given compound, integer.
BondChiralUndefCount [BCUC, BCUCNT]: Total count of undefined
chiral bonds in a given compound, integer.
CompleteSynonym [CSYN, CSYNO]: Compound's synonyms, based on all substance related to this compound.
Complexity [CPLX]: Compound complexity.
CompoundID [CID]: Compound ID. Same as [UID].
CovalentUnitCount [CUC, CUCNT]: Integer.
CreateDate: Date this compound created in PubChem.
Element [ELMT, EL]: Chemical element in a compound.
ExactMass [EMAS, EXMASS]: The calculated mass of an ion or a molecule
containing most likely isotopic composition for a single random molecule,
corresponding to mass of most intense mol/molecule peak in a MS spec. A real number.
HeavyAtomCount [HAC, HACNT]: Atom count in a compound except hydrogen,
integer.
HydrogenBondAcceptorCount [HBAC, HBACNT]: Hydrogen bond acceptors for a
compound, integer.
HydrogenBondDonorCount [HBDC, HBDCNT]: Hydrogen bond donors for a
compound, integer.
InactiveAid [IA]: Inactive BioAssay identifier, integer. This choice is intended for users querying for inactive
compounds for a particular assay. BioAssay ID (AID) should be provided.
InactiveAidCount [IC, ICNT]: Using this filter users can query for compounds which are inactive in a certain number of assays
InChI [INCH, INCHI]: IUPAC International Chemical Identifier.
InChI string can be searched through the Entrez PubChem databases.
e.g.To search with the InChI string of aspirin: "InChI=1/C9H8O4/c1-6(10)13-8-5-3-2-4-7(8)9(11)12/h2-5H,1H3,(H,11,12)/f/h11H":
Type or paste "InChI=1/C9H8O4/c1-6(10)13-8-5-3-2-4-7(8)9(11)12/h2-5H,1H3,(H,11,12)/f/h11H"[InChI]
into the
PubChem Compound, or
PubChem Substance, or the
Entrez Global search box, then click Go button.
Note:
The quote marks and the square brackets are
required.
'InChI=' is required.
|
IsotopeAtomCount [IAC, IACNT]: Isotope atom numbers in a compound.
IUPACName [UPAC, IUPAC]: Standard IUPAC name for compound.
MeSHDescription [MHD]
MeSHTerm [MSHT, MESHT]: Medical Subject Heading term.
Note that MeSH entry terms (synonyms for the Medical Subject Heading term) are also indexed.
MeSHTreeNode [MSHN, MESHTN]: Medical Subject Heading tree node (tree structures). Searching for a term corresponding to a higher level node in the MeSH hierarchy will find records matching any MeSH terms beneath that node. For example, "Penicillins[MeSHTreeNode]" will find records linked to MeSH terms "Oxacillin", "Cloxacillin", etc.
Note that entry terms (synonyms) for MeSH tree nodes are also indexed.
MolecularWeight [MW, MWT, MOLWT]: Mass of a molecule calculated
using the average mass of each element weighted for its natural isotopic
abundance. E.g., Carbon has two natural isotopes 12 and 13 with relative
abundances of 98.9% and 1.1% to yield an average mass of 12.011 g/mol. A real
number.
MonoisotopicMass [MMAS, MIMASS]: Mass of a molecule calculated
using the mass of the most abundant isotope of each element. E.g., Carbon has a monoisotopic mass of 12.000 g/mol. A real number.
PharmAction [PHMA, PHARMA]: MeSH pharmacological actions.
RotatableBondCount [RBC, RBCNT]: Count of rotatable bonds
SourceID [SRID, SRCID]: Depositor's external id.
SourceName [SRC, SRCNAM, SRCNAME]: Depositor name officially recorded
in PubChem databases. For current data sources look here
SourceCategory [SRCC, SRCCAT, SRCCATG]: Depositor categories. For more information and possible categories
look here
SubstanceID [SID]: Substance identifier, integer.
Synonym [SYNO]: Synonyms for substance.
TautomerCount [TC, TCNT, TTMC]: Possible tautomer count for each given
structure, no more than 200, integer.
TestedConcentration : Using this filter users can query for compounds which are tested under certain concentrations(uM) in biological assays.
TotalAidCount [TAC]: TotalAidCount includes any assay that a compound is tested, it should cover
active/inactive/inconclusive/unspecified
TotalFormalCharge [TFC, CHG, CHRG]: Total formal charge.
TPSA[TPSA]: Topological Polar Surface Area.
XLogP [XLGP, LOGP]
PubChem Substance:
All [ALL]: All of the following fields are searched. If a string query is presented without
a field alias, by default, [ALL] is searched.
Uid [UID]: The integer represents SID for Pcsubstance
database. By default, an integer without a field alias is recognized as a UID. Same as [SID].
Filter [Filter]: Limits the records. A number of filters are available to restrict the search to substances
with particular information. The specialized Filters in this database are:
- has_mesh: records with associated MeSH terms
- has_pharm: records with associated pharmacological actions
- has_parent: records that have a parent structure
- has_no_parent: records that do not have a parent
ActiveAid [AA]: Active BioAssay identifier, integer. This choice is intended for users querying for active
substances for a particular assay.
ActiveAidCount [AC, ACNT]: Using this filter users can query for compounds which are active in a certain number of assays.
ActiveAidRatio [AAR]: Ratio should be between zero and 1. Ratio equals to the number of
BioAssays where substances were tested active divided by number of BioAssays where substances
tested with any result.
(# BioAssays where tested active) / (# BioAssays tested with any result)
ActiveConcentration: Using this filter users can query for substances which show certain extent of active concentration(uM) in biological assays.
AssaySourceName [ASRC, ASRCNAM, ASRCNAME]: Allows filtering of by assay source name.
For available data sources look here
AtomChiralCount [ACC, ACCNT]: Total count of chiral atoms in a given
compound, integer.
AtomChiralDefCount [ACDC, ACDCNT] -- Total count of defined chiral
atoms in a given compound, integer.
AtomChiralUndefCount [ACUC, ACUCNT] -- Total count of undefined
chiral atoms in a given compound, integer.
BioAssayID [BAID, AID]: BioAssay identifier, integer.
BondChiralCount [BCC, BCCNT]: Total count of chiral bonds in a given
compound, integer.
BondChiralDefCount [BCDC, BCDCNT]: Total count of defined chiral
bonds in a given compound, integer.
BondChiralUndefCount [BCUC, BCUCNT]: Total count of undefined
chiral bonds in a given compound, integer.
Comment [CMT]: Substance or BioAssay comment.
CompleteSynonym [CSYN, CSYNO]: Compound's synonyms, based on all substance related to this compound.
Complexity [CPLX]: Substance structure complexity.
ComponentCID [CCID]: Component compound identifier.
CompoundID [CID]: Compound identifier, integer.
CovalentUnitCount [CUC, CUCNT]: Integer.
DepositDate [DDAT, DEPDAT]: Deposition timestamp for a substance.
Element [ELMT, EL]: Chemical element in a substance/compound.
ExactMass [EMAS, EXMASS]-- The calculated mass of an ion or a molecule
containing most likely isotopic composition for a single random molecule,
corresponding to mass of most intense ion/molecule peak in a MS spec. A real number.
HeavyAtomCount [HAC, HACNT]: Atom count in a compound except hydrogen,
integer.
HydrogenBondAcceptorCount [HBAC, HBACNT]: Hydrogen bond acceptors for a
compound, integer.
HydrogenBondDonorCount [HBDC, HBDCNT]: Hydrogen bond donors for a
compound, integer.
InactiveAid [IA]: Inactive BioAssay identifier, integer. This choice is intended for users querying for inactive substances for a particular assay.
BioAssay ID (AID) should be provided.
InactiveAidCount [IC, ICNT]: Using this filter users can query for substances which are inactive in a certain number of assays
InChI [inchi]: IUPAC International Chemical Identifier.
InChI string can be searched through the Entrez PubChem databases.
e.g.To search with the InChI string of aspirin:
"InChI=1/C9H8O4/c1-6(10)13-8-5-3-2-4-7(8)9(11)12/h2-5H,1H3,(H,11,12)/f/h11H":
Type or paste "InChI=1/C9H8O4/c1-6(10)13-8-5-3-2-4-7(8)9(11)12/h2-5H,1H3,(H,11,12)/f/h11H"[InChI]
into the
PubChem Compound, or
PubChem Substance, or the
Entrez Global
search box, then click Go button.
Note:
The quote marks and the square brackets are
required.
'InChI=' is required.
|
IsotopeAtomCount [IAC, IACNT]: Isotope atom numbers in a compound.
IUPACName [UPAC, IUPAC]: Standard IUPAC name for compound.
MeSHDescription [MHD]
MeSHTerm [MSHT, MESHT]: Medical Subject Heading term.
Note that MeSH entry terms (synonyms for the Medical Subject Heading term) are also indexed.
MeSHTreeNode [MSHN, MESHTN]: Medical Subject Heading tree node (tree structures). Searching for a term corresponding to a higher level node in the MeSH hierarchy will find records matching any MeSH terms beneath that node. For example, "Penicillins[MeSHTreeNode]" will find records linked to MeSH terms "Oxacillin", "Cloxacillin", etc.
Note that entry terms (synonyms) for MeSH tree nodes are also indexed.
ModifiedDate: Date this substance record is modified.
MolecularWeight [MW, MWT, MOLWT]: Mass of a molecule calculated
using the average mass of each element weighted for its natural isotopic
abundance. E.g., Carbon has two natural isotopes 12 and 13 with relative
abundances of 98.9% and 1.1% to yield an average mass of 12.011 g/mol. A real
number.
MonoisotopicMass [MMAS, MIMASS]: Mass of a molecule calculated
using the mass of the most abundant isotope of each element. E.g., Carbon has a
monoisotopic mass of 12.000 g/mol. A real number.
PharmAction [PHMA, PHARMA]: MeSH pharmacological actions.
RotatableBondCount [RBC, RBCNT]: Count of rotatable bonds
SourceCategory [SRCC, SRCCAT, SRCCATG]: Depositor categories. For more information and possible
categories look here
SourceID [SRID, SRCID]: Depositor's external id.
SourceName [SRC, SRCNAM, SRCNAME]: Depositor name officially recorded
in PubChem databases. For current data sources look here
SourceReleaseDate [SRD, SRDAT, RLSDAT]
StandardizedCID [SCID]: Standardized compound identifier, integer.
SubstanceID [SID]: Substance ID. Same as [UID].
Synonym [SYNO]: Synonyms for substance.
TautomerCount [TC, TCNT, TTMC]: Possible tautomer count for each given
structure, no more than 200, integer.
TestedConcentration : Using this filter users can query for substances which are tested under certain concentrations(uM) in biological assays.
TotalAidCount [TAC]
TotalFormalCharge [TFC, CHG, CHRG]: Total formula charge.
TPSA [TPSA]: Topological Polar Surface Area.
XLogP [XLGP, LOGP]
PubChem BioAssay:
All [ALL]: All of the following fields are searched. If a string query is presented without
a field alias, by default, [ALL] is searched.
Uid [UID]: The integer represents AID for Pcassay database. By default, an integer
without a field alias is recognized as a UID.
Filter [Filter]: Limits the records. A number of filters are available, to retrieve
records in the same or other databases that the current BioAssay records are cross-referenced
to.
ActiveCidCount [ACC, ACCNT]: Number of unique chemicals (identified by CID--compound
identifiers from Pccompound) that are considered as active in a BioAssay.
ActiveSidCount [AC, ACNT]: Number of substances (identified by SID--substance
identifier from Pcsubstance) that are considered as active in a BioAssay.
Activity Outcome Method: Description on how activity outcome is determined.
Choices of search query include:
-
screening: reports number of 'Screening' assay
- Single Concentration Activity Observed: Activity outcome was
defined based on the percentage of inhibition from test at a single dose
-
confirmatory: reports number of 'Confirmatory' assay
- Concentration-Response Relationship Observed: Activity outcome
was defined based on EC50/IC50 values and so forth, derived from dose
response curves following tests with multiple concentrations
-
summary: reports number of 'Summary' assay
- Candidate Probes/Leads with Supporting Evidence: An assay which
summarizes information from multiple assays
-
other: reports number of assays in the 'Other' category
- An assay which does not fall into the above categories
AssayComment [AssayComment]: Comment for the BioAssay.
AssayDescription [ADES, ADESC, ADSC]: Description for the BioAssay.
AssayName [ANAM, ANAME]: Name of a BioAssay provided by depositor.
AssayProtocol [APRL, APRTL]: Protocol for a BioAssay provided by depositor.
AssayComment [ACMT, ACMMNT]: comment for a BioAssay provided by depositor.
CompoundIDActive [CIDA]: CID(compound identifier from Pccompound) of a chemical
that is considered as active in a BioAssay.
CompoundIDTested [CIDT]: CID(compound identifier from Pccompound) of a chemical that is tested in a BioAssay.
InactiveCidCount [IACC, IACCNT]: Number of unique chemicals (identified by CID--compound identifiers from Pccompound) that are considered as inactive in a BioAssay.
InactiveSidCount [IAC, IACNT]: Number of substances (identified by SID--substance identifier from Pcsubstance) that are considered as inactive in a BioAssay.
InconclusiveCidCount [ICC, ICCNT]: Number of unique chemicals (identified by CID--compound identifiers from Pccompound) that are considered as inconclusive regarding to be active or inactive in a BioAssay.
InconclusiveSidCount [IC, ICNT]: Number of substances (identified by SID--substance identifier from Pcsubstance) that are considered as inconclusive regarding to be active or inactive in a BioAssay.
MeSHDescriptionActive MHDA]: Medical Subject Heading descriptions that are associated with an active chemical structure in a BioAssay.
MeSHDescriptionTested [MHDT]: Medical Subject Heading descriptions that are associated with any chemical structure tested in a BioAssay.
MeSHTermActive [MHTA]: Medical Subject Heading terms that are only associated with an active chemical structure in a BioAssay.
MeSHTermTested [MHTT]: Medical Subject Heading terms that are associated with any chemical structure tested in a BioAssay.
ModifiedDate [MDAT, MDATE]: Last date when a BioAssay data content is modified. Date format is yyyy/mm/dd. mm and dd are optional.
PharmActionActive [PHAA]: MeSH pharmacological actions that are associated with an active chemical structure in a assay.
PharmActionTested [PHAT]: MeSH pharmacological actions that are associated with an any chemical structure tested in a assay.
ReadoutCount [RC, RCNT]: Number of total test result fields(readouts) in a BioAssay
ReleaseDate [RDAT, RDATE]: Date when a BioAssay data is released to public by PubChem. Date format is yyyy/mm/dd. mm and dd are optional.
SourceCategory [SRCC, SRCCAT, SRCCATG]: Category of BioAssay data source
SourceName [SNME, SNAME]: Source name of a BioAssay data specified by depositor.
SubstanceIDActive [SIDA]: SID(substance identifier from Pcsubstance) of a substance that is considered as active in a BioAssay.
SubstanceIDTested [SIDT]: SID(substance identifier from Pcsubstance) of a substance that is tested in a BioAssay.
SynonymActive: MESH names and synonyms that are associated only with an active chemical structure in a BioAssay.
SynonymTested [SYNT]: MESH names and synonyms that are associated with any chemical structure tested in a BioAssay.
TidDescription [TDES, TIDD, TDESC, TDSC]: Description of a test result field in a BioAssay
TidName [TNAM, TIDN, TNAME]: Name of a test result field in a BioAssay
TestedConcentration : Using this filter users can query for assays where substances are tested in certain concentration(uM).
TotalCidCount [TCC]: Total number of unique chemicals (identified by CID--compound identifiers from Pccompound) that are tested in a BioAssay.
TotalSidCount [TSC]: Total number of substances tested in a BioAssay.
XRefAid [XRAD]: NCBI Entrez PubChem BioAssays identifier.
XRefAsurl [XRAL]: URL of the BioAssay provided by depositor.
XRefComment [XRCT]
XRefDburl [XRDL]: URL of Depositor's organization.
XRefGene: NCBI Entrez Gene identifier referred by a BioAssay.
XRefGi [XRGI]: NCBI Entrez Protein/Nucleotide GI number referred by a BioAssay.
XRefMmdb [XRMB]: NCBI Entrez Molecular Modeling Database(MMDB) identifiers(MMDB ID)
referred by a BioAssay.
XRefNucleotidegi [XRefNCGI, XRNI]: NCBI Entrez Nucleotide identifier
referred by a BioAssay.
XRefOmim [XRMM]: NCBI Entrez OMIM identifier.
XRefPmid [XRPD]: NCBI Entrez PubMed identifiers(PubMed ID) referred by a BioAssay.
XRefProteingi [XRefPRGI, XRPI]: NCBI Entrez Protein identifier referred by a BioAssay.
XRefSburl [XRSL]
XRefTaxonomy [XRTY]: NCBI Entrez Taxonomy identifier.
| PubChem Batch Processing Utility |

|
PubChem batch processing utility allows power users to
process PubChem data in batch mode. Please follow the procedures provided to
perform your job. Note: Please limit your batch to 50,000 records as this is the
limit for the PubChem download facility.
Utility Type:
- Please choose one utility that you want to process.
Hints to generate an ID-Map file:
- Do any search at the Entrez PubChem Substance database, e.g. searching term "Cu[el]"
- At the Display dropdown list, choose "ID Map"
- At the "Send to" dropdown list, choose "File"
- Save the file for the PubChem Batch Utility use.
Submit Your Job:
- Once you finish step 1 and 2, click the Go button to submit your job.
New Job:
- To start a new job, click this link.
Q: What is PubChem ?
A: PubChem is a component of NIH's
Molecular Libraries
Roadmap Initiative. It provides
information on the biological activities of small molecules. PubChem is
organized as three linked databases within the NCBI's
Entrez information
retrieval system. These are
PubChem Substance,
PubChem Compound, and
PubChem BioAssay. PubChem also provides a fast chemical
similarity search tool.
Q: What is PubChem Substance ?
A: PubChem Substance records contain substance information
electronically submitted to PubChem by depositors. This includes any chemical
structure information submitted, as well as chemical names, comments, and links
to the depositor's web site.
Q: What is PubChem Compound ?
A: PubChem compound records
comprise a non-redundant set of standardized and validated chemical structures.
A compound record may link to more than one PubChem Substance record, if
different depositors supplied the same structure. Chemical names shown in
PubChem Compound records are a composite derived from all linked substances,
with default ranking of names by weighted frequency of use.
Q: What does the depositor's category tell users and what are the existing depositor categories ?
A:
The depositor categories indicate the type of information one may expect to find when following the
depositor substance URL or the type of information provided by the depositor.
A list of possible categories include the following:
| Status |
Meaning |
| Biological Properties |
Depositor provides information about the biological properties of a substance or compound |
| Chemical Reactions |
Depositor provides information about the reactivity, synthesis, or known reactions of a substance or compound |
| Imaging Agents |
Depositor provides information about the contrast agent or imaging agent used in, for example, MRI's |
| Journal Publishers |
Depositor is a journal publisher and has articles published about a substance or compound |
| Metabolic Pathways |
Depositor provides information on the metabolic pathways involving a substance or compound |
| Molecular Libraries Screening Center Network |
Depositor is part of the NIH Molecular Libraries Screening Center Network (MLSCN) |
| NIH Substance Repository |
Depositor is an NIH Molecular Libraries Small Molecule Repository servicing the MLSCN |
| Physical Properties |
Depositor provides information about the experimental physical properties of a substance or compound |
| Protein 3D Structures |
Depositor provides information about the experimental 3-D structure of a substance or compound |
| Substance Vendors |
Depositor is a seller of a substance or compound |
| Theoretical Properties |
Depositor provides information about the theoretical properties of a substance or compound |
| Toxicology |
Depositor provides information about the toxicological properties of a substance or compound |
Q: Why search PubChem Substance and/or Compound ?
A: It is useful to search PubChem's
Substance database when one is looking for information from a particular
depositor exclusively, and/or when one is looking for information on substances
such as natural product extracts which may not have associated chemical
structure information. These special cases aside, it is generally most useful to
search for chemical names or structures in PubChem's Compound database. This
provides a concise view, combining information derived from multiple Substance
records that specified the same structure. PubChem's structure search service
operates on PubChem's Compound database exclusively.
Q: What is PubChem BioAssay ?
A: The PubChem BioAssay Database
contains BioActivity screens of chemical substances described in
PubChem
Substance. It provides searchable descriptions of each BioAssay,
including descriptions of the conditions and readouts specific to that screening
procedure.
Q: How does PubChem assign Substance identifiers ?
When a substance is revoked by the depositor, can I still see the old record ?
A: A PubChem Substance SID is assigned to each unique external registry ID provided by
a PubChem data source. A depositor may "revoke" (or otherwise deprecate) a PubChem SID at any time
for any reason. However, the link to the "revoked" PubChem SID lives on in perpetuity.
There will be a message stating the depositor deprecated the SID, but the link to the archived
information will still be available. In addition, the PubChem CID's pointed to by the old version
of a PubChem SID at the time it was versioned or deprecated will also be available.
Q: How does PubChem assign Compound identifiers ?
Will the structure represented by a CID ever change ?
A: A PubChem Compound CID is assigned to each unique chemical structure.
It is possible that different tautomeric forms of the same compound to have different CID's.
The chemical structure represented by a CID is permanent. The URL links to the compound summaries
are stable (always live), regardless if any (or no) substance points to them.
Q: How does PubChem process my deposited structures ?
A: The conversion of the deposited information goes through a series of validation steps (to confirm the structure is "valid") and then a series of standardization/normalization steps to remove VB redundancy.
The validation steps consist of:
Atom verification: do all atoms correspond to a known atomic element? E.g., "*" is not a known atom
Implicit hydrogens are assigned to organic elements using simple valence rules, e.g., methane "C" gets four implicit hydrogens assigned to it.
Functional group standardization: common incorrect and hypervalent representations of functional groups are "fixed", e.g., nitro groups represented by N(=O)=O become [N+](=O)[O-]
Atom valences are validated: do all atoms have an "allowed" valence? E.g., five bonds to carbon is not valid
The standardization steps consist of:
Valence bond (VB) canonicalization: equivalent/alternate VB/tautomeric forms of a structure are normalized into a single representation
Aromaticity detection: structure aromaticity is detected and validated to be kekulizable
StereoChemistry detection: SP3 and SP2 stereo centers are detected and stereo-wedge placement standardized
Explicit hydrogen assignment: implicit hydrogens are converted to be explicit
Subsequent additional processing includes 2D coordinate layout assignment.
Q: How do I process a text search with PubChem
databases ?
A: PubChem's Substance, Compound, and BioAssay databases are fully
integrated within NCBI's Entrez data retrieval system. You can process any name,
keyword, or ID search through the Entrez system. The
PubChem homepage also provides a
search box. For a specific database query, see related content in the help
document above.
Q: How do I perform a structure search ?
A: You can perform a structure search through the PubChem structure database.
PubChem provides two search interfaces, basic structure search and advanced structure
search. For more information, visit
structure
search help.
Q: How do I save my search result ?
A: To save your search from Entrez, you can use either the
PubChem download facility or the
Entrez generic search-save tool. You can get more help by clicking the two links
above.
Q: Sometime I see errors in the substance record,
where I should report ?
A: PubChem doesn't have curators and never changes/edits substance records.
They remain as supplied by our depositors, just as with GenBank records. You can
follow PubChem substance summary page to find the original record from
the depositor's page, and report the error. Once the error is corrected by
the depositor, PubChem will implement it at next update. For compound
property/descriptor errors, you can report to the
NCBI help desk.
Q: What are exact mass and monoisotopic mass for
a substance/compound ?
A: An exact mass is the most likely isotopic composition for a single random molecule,
corresponding to mass of most intense ion/molecule peak in a mass spectrum.
A monoisotopic mass is a molecule calculated using the mass of the most abundant isotope of each element.
For example, carbon has a monoisotopic mass of 12.000 g/mol.
Exact mass and monoisotopic mass are the same for more than 90% of structures but differ when atom counts are
such that presence of one or more lower abundance isotopes is most probable.
For example, carbon tetrachloride, CCl4, PubChem CID 5943,
has an exact mass of 153.872, where,
in this case, the prototypical compound is made of three 35Cl, one
37Cl, and one 12C.
The monoisotopic mass for carbon tetrachloride is 151.875, where in this case all chlorine atoms are assumed to be
35Cl,
with isotope abundance of 75.77%, and the carbon atom is assumed to be 12C, with isotope abundance of 98.9%.
In many cases, these two masses are identical, except for compounds with four or more Cl atoms, two or more Br atoms,
or other elements not dominated by a single isotope, or for really large compounds such as with the number of carbons
greater than 99, i.e., for C100 the exact mass will be 12C * 99 +
13C * 1,
while the monoisotopic mass will be 12C * 100.
Q: How do I find INCHI version and parameters?
A:
The InChI version and parameters are detailed in the ASN.1 and XML data
for each compound. For example, for aspirin, you can find the information you
are seeking by viewing the ASN.1 record:
http://pubchem.ncbi.nlm.nih.gov/summary/summary.cgi?cid=2244&disopt=DisplayASN1
If you scroll down to the InChI property record, you will find:
{
urn {
label "InChI",
datatype string,
parameters "options {auxnonr donotaddh w0 fixedh recmet newps}",
implementation "E_INCHI",
version "1.0.1",
software "InChI",
source "nist.gov",
release "2007.09.04"
},
value sval "InChI=1/C9H8O4/c1-6(10)13-8-5-3-2-4-7(8)9(11)12/h2-5H,1H3,(H,11,12)/f/h11H"
},
The InChI version is (currently) 1.0.1. We are using the parameter
options "auxnonr donotaddh w0 fixedh recmet newps".
NCBI provides PubChem training courses:
AID: PubChem's BioAssay (protocol) identifier, a non-zero integer.
CID: PubChem's compound identifier, a non-zero integer for a unique chemical structure.
Complexity : The complexity rating of the compounds is a rough estimate of
how complicated a structure is, seen from both the point of view of the elements contained and the displayed
structural features including symmetry. However, neither stereochemistry nor isotope labeling are used as
auxiliary criteria. The value is computed using the Bertz/Hendrickson/Ihlenfeldt formula. A scaling factor
for aromaticity is used so that the complexity of benzene is the same as of cyclohexane.
It is a floating point value, ranging from 0 (simple ions) to several thousand (complex natural products).
Generally larger compounds are more complex than smaller ones, but highly symmetrical compounds,
or compounds with few distinct atom types or elements are downgraded. Complexity is only loosely
correlated with synthetic accessibility.
The most complex compound in PubChem is CID
6338588
(C124H185N9O207S36) with a complexity rating of about 18425. The average complexity of the
structures in PubChem compound database is about 551.
Comments: List all depositor's comments and additional information
for this substance.
Component: For mixture substance/compound, component is one of the single molecule.
Compound: Chemical representatives in substances. Chemical structure presented in a compound is standardized
through PubChem's data pipeline. A mixture substance may have several standardized compounds. A compound record is structurally
unique in the PubChem compound database.
Computed Descriptors: Information to describe the compound in different formats, including SMILES,
InChI, IUPAC names.
Computed Properties: These data are calculated from the compound, including molecular weight, formula, XLogP, etc.
Depositors Category: Depositors category tells users that there is an additional
category-specific information either on depositors substance summary page or on the depositor's web-site.
Deprecated Compound: A Compound CID which
has no links to any substance. This may occur as PubChem modifies processing. A
deprecated compound will not be available within Entrez.
HBA: Number of hydrogen acceptors in the structure. Classification of hydrogens follows [J. Chem. Inf. Comput. Sci. 1997,37, 615-621].
HBD: Number of hydrogen donors in the structure. Classification of hydrogens
follows [J. Chem. Inf. Comput. Sci. 1997,37, 615-621].
Heavy Atom: All atoms except hydrogen.
InChI: IUPAC International Chemical Identifier.
Learn more... InChI string
can be searched through the Entrez PubChem databases. Click
here to see the example. Old Version Substance --
Substance versions are considered to be "old" when a more recent update is
provided by the depositor.
Molecular Formula: A way of expressing information about the atoms that constitute a particular chemical molecule.
Molecular Weight: The
molecular weight is the sum of all atomic weights of the constituent atoms in a compound, measured in gr/mol. In the absence of explicit isotope labeling, averaged natural abundance (which may, for example in case of Li and U compounds, not be identical to purchasable material) is assumed. If an atom bears an explicit isotope label, 100% isotopic purity is assumed at this location, even for short-lived radioactive isotopes where this is often physically unrealistic. At this moment, it is not possible to deposit more detailed isotope composition information into the PubChem database. Pseudo-atoms which are not an element have an atomic weight of 0 g/mol.
Revoked BioAssay: When a depositor removes an
assay that the depositor previously deposited into PubChem, the assay is considered revoked.
A revoked assay will not be available within Entrez..
Revoked Substance: When a depositor removes a
substance from their substance collection, the substance is considered revoked.
A revoked substance will not be available within Entrez.
SID: PubChem's substance identifier, a non-zero integer for a deposited substance.
SMILES: Simplified Molecular Input Line Entry System,
a line notation (a typographical method using printable characters) for
entering and representing molecules.
Learn
more..
You can also find more related information form PubChem's document section in
PDF or
Text.
SMARTS: A language that allows you to specify substructures using rules
that are straightforward extensions of SMILES.
Learn more..
Substance: Individual record object collected from depositors, representing a sample used at BioAssay.
Substance Category: Substance categories (one or more) are
assigned to each depositor, based on nature of that depositor's institution and the type of data they supply.
Suppressed Compound: A Compound CID that
links only to an old version substance. A suppressed compound will not be
available within Entrez.
Synonyms: All names, trivial names, synonyms, frequently used IDs, and other names collected from depositors. In
the compound summary page, synonyms are
distinct synonyms from all corresponding substances.
TPSA --
Topological Polar Surface Area. This is an estimate of the area (in Å squared) which is polar. The implementation follows [J. Med. Chem.
2000, 43, 3714-3717.]. It is a simple method - only N and O are considered, 3D coordinates are not used, and there are various precomputed factors for different hybridizations, charges and participation in aromatic systems.
Version: PubChem substance version number is
incremented when an update is provided by the depositor.
Xref: The external references/links to PubChem database records.
XLogP: A partition coefficient or distribution coefficient
that is
a measure of differential solubility of a compound in two solvents.
Learn more..
From November 2006, the PubChem uses version 2 of the algorithm of the reference [
Perspectives in Drug Discovery and Design. 2000,
19, 47-66.] to generate the XlogP value.
|



