Random Intensity Datasets: 60 Cases, 60 Controls
Brian T. Luke (lukeb@ncifcrf.gov)Return to Contents
These five pairs of datasets contain 300 features with 60 Cases and 60 Controls. These datasets are constructed with random peak intensities so that they contain no biological information. Structure of the Datasets contains a general description of datasets that can be used by programs within the BioMarker Development Kit (BMDK). Since the Cases and Controls are stored in different files, the class indices are not included in the data. Each feature has a single label, but they are simply "F-00001" through "F-00300". Each dataset has an associated document that describes the results of an analysis using the BioMarker Development Kit (BMDK), and classifiers based on a decision tree (DT) and a medoid classification algorithm (MCA). To reduce the amount of repeated information in these tables of results, Description of the Tables gives details about each table.
| Analysis | #Cases #Controls |
#Features | Case Dataset |
Control Dataset |
Analysis |
| Random_Intensity_60_1a | 60 | 300 | case_60_1a.txt | control_60_1a.txt | Tables |
| Random_Intensity_60_2a | 60 | 300 | case_60_2a.txt | control_60_2a.txt | Tables |
| Random_Intensity_60_3a | 60 | 300 | case_60_3a.txt | control_60_3a.txt | Tables |
| Random_Intensity_60_4a | 60 | 300 | case_60_4a.txt | control_60_4a.txt | Tables |
| Random_Intensity_60_5a | 60 | 300 | case_60_5a.txt | control_60_5a.txt | Tables |
The following table lists the best classification observed results for each dataset-pair.
| Set | NPB | BMDK-1 | BMDK-2 | BMDK-3 | DT | MCA-5 | MCA-6 | MCA-7 |
| 60_1a | 22 | 136.7 | 138.3 | 140.1 | 176.7 | 191.7 | 193.3 | 195.0 |
| 60_2a | 29 | 121.7 | 129.7 | 128.9 | 176.7 | 193.3 | 193.3 | 195.0 |
| 60_3a | 26 | 133.3 | 140.1 | 133.6 | 178.3 | 193.3 | 193.3 | 193.3 |
| 60_4a | 22 | 130.0 | 139.3 | 133.7 | 176.7 | 193.3 | 193.3 | 195.0 |
| 60_5a | 27 | 151.7 | 137.0 | 136.2 | 171.7 | 193.3 | 193.3 | 195.0 |
For each set of Cases and Controls, BMDK uses 10 different methods to search for putative biomarkers, and the number of putative biomarkers (NPB) identified for each set is listed in the second column (the Tables shown in the links above give details on which procedures selected which features). BMDK only uses these putative biomarkers to construct the final classifier based on a distance-dependent K-nearest neighbor algorithm. This classifier allows for an "undetermined" classification, so the quality metrics shown above are the sum of the overall sensitivity and specificity minus the percent "undetermined" from a leave-one-out cross-validation analysis, with the constraint that no more than 5% of the samples can be "undetermined". The third, fourth and fifth columns list the best result using between one and three of the putative biomarkers, respectively. For the DT and MCA classifiers, the quality is the sum of the sensitivity and specificity.
With the exception of the 1-feature classifier for Set 60_5a, none of the final BMDK classifiers produced a sensitivity and specificity above 70%. The 2-feature classifier for set 60_3a yielded a sensitivity of 69.6% and a specificity of 74.6% with five samples (4.2%) receiving an undetermined classification. The 3-feature classifier for Set 60_1a also had a sensitivity of 69.6%, a specificity of 74.6% with a 4.2% of the samples (five samples) receiving an "undetermined" classification. The 1-feature classifier for Set 60_5a yielded an relatively high quality score (sensitivity=78.3%, specificity=73.3%, and no undetermined samples). The intensities of this feature are shown in the following figure for the Cases (left column) and Controls (right column).
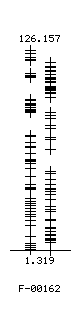
This feature produced these relatively good results because several of the Cases had intensity in regions containing virtually no Controls and several Controls in regions with very few cases. Since a random distribution of intensities is not uniform for a finite number of samples, it is possible to obtain a relatively good result by chance [Ran-05a, Ran-05b]. A visual inspection of the peak intensities is therefore necessary, since this feature does not have a sufficient difference in the ranges of intensities for Cases and Controls to represent a true biomarker.
The best DT classifiers (Column 6) containing up to seven decision nodes misclassified between four and seven samples, yielding an average sensitivity and specificity of at least 85.8%. For four of the five datasets, the average sensitivity and specificity varied between 88.3 and 89.1% across all 120 samples. The final three columns of the preceding table show the best results for an MCA classifier using five, six, and seven features, respectively. The best 5-feature classifiers had an average sensitivity and specificity of at least 95.8%%, while the 6- and 7-feature classifiers has an average sensitivity and specificity of at least 96.6%. In all cases, the MCA classifier was constructed after effectively separating the data into a training set containing 40 Cases and 40 Controls, and a testing set containing 20 Cases and 20 Controls.
It is clear that the fingerprint-based methods are able to classify these samples to a very high accuracy, even though these datasets are constructed to contain no biological information.
(Last updated 9/1/07)




