Liver
Library
Creating
a Microarray for Hepatotoxicants
Mechanical improvements in high-throughput applications continue to increase
the utility of the microarray approach for investigating toxic effects on genes.
But improvements in the content of arrays may be the key to maximizing the value
of these technologies, according to a paper in this month's issue [EHP
111:863–870]. In the report, researchers at Abbott Laboratories
and Rosetta Inpharmatics, led by senior research scientist Jeffrey Waring, lay
out the development of a microarray specifically constructed for studying the
effects of hepatotoxicants.

image credit: Matt Ray/EHP, Joseph Tart/EHP, Artville, EyeWire
|
| Array of hope. Scientists
have a new tool for understanding liver responses to toxic exposures. |
Work in toxicogenomics has so far focused primarily on hepatotoxicity because
of the importance of the liver as a site of toxic response. Whereas earlier
toxicology-focused arrays were put together using DNA libraries from normal
or diseased tissues, building a library from toxicant-challenged animals is
a new approach. Because these animals were specifically expressing genes regulated
in response to toxic exposure, it was possible for the Abbott-Rosetta team
to enrich for genes regulated by toxic compounds, making their array a highly
specific tool for understanding the function of rat liver undergoing toxic exposure.
Understanding how gene expression changes when animals face different toxicants
is especially important in light of growing evidence suggesting that even dissimilar
toxicants can elicit similar response mechanisms calling similar groups of genes
into play.
The array was made from cDNA derived from RNA from male Sprague-Dawley rats
exposed to 52 different compounds at two levels during three-day toxicity studies.
Applying the compounds orally, interperitoneally, or intravenously (depending
on the compound), the scientists exposed 3 rats to both levels of each toxicant.
They formed the pool of RNAs used to make the array from a total of 312 exposed
rats. The exposure compounds induce a variety of toxic mechanisms including
DNA damage, cirrhosis, oxidative stress, steatosis (accumulation of fat in the
liver), and necrosis.
The scientists enriched their library for genes induced by exposure to the
study toxicants by using a subtractive hybridization approach that allowed them
to eliminate transcripts that were also present in nonexposed animals. Using
animals exposed for 3 days allowed induction of gene-level responses in the
liver, but avoided capturing genes involved in the later processes of secondary
inflammation or fibrosis. Sequencing clones from the library allowed identification
of more than 2,700 expressed putative genes. About 20% of these genes, the scientists
indicate, do not appear to have been previously described.
Genes from this library make up about 25% of the array, which contains 25,000
probes. The other 75% includes rat genes with known human orthologs (which help
compare gene expression patterns between species), genes allowing comparisons
between specific and nonspecific hybridization, hybridization targets to allow
comparisons of hybridization intensity, and other controls.
The researchers say these gene expression profiles can be used to build a
predictive database encapsulating biological responses to toxic insult. If the
concept of "guilt by association" is to prove accurate, they write--if compounds
are considered to have toxic liabilities when they closely associate with a
known toxicant--it is extremely important to equip the array with the correct
genes to distinguish the mechanism of toxicity.
Victoria McGovern
Monitoring
Estrogenic Effects
A
Genomics Approach
Genomics, the revolutionary field that promises to one day reveal the genetic
code of every living organism, is opening up unforeseen opportunities for advances
in many areas of the life sciences. In this issue, a team led by Patrick Larkin
of the University of Florida in Gainesville and EcoArray LLC describes a genomics
approach to monitoring toxic chemicals in the environment and uncovering their
effects on organisms at the molecular level [EHP
111:839-846].
Larkin and colleagues hope to produce an easy-to-use biomarker test capable
of detecting metabolic pathways affected by environmental chemicals, and ultimately
to formulate specific gene profiles that will permit identification of particular
chemical contaminant exposures. In this article, the team describes an expression
profiling model system for endocrine-disrupting compounds (EDCs) that mimic
estrogens.
Natural and synthetic estrogens are found in pharmaceuticals, industrial by-products,
and pesticides, and can cause human health effects including vaginal cancer
and reproductive tract abnormalities. Because estrogen is a female reproductive
hormone, genes in the estrogen pathway are normally not highly expressed in
males. However, when male fish are exposed to natural or synthetic estrogens,
the result is an increase in the expression of female-specific genes. The estrogen
pathway has been highly conserved during vertebrate evolution--it is shared
by many different organisms--so changes due to exposure in fish may presage
effects in other animals, including humans.
The team created a gene array by cloning 30 genes--some involved in the estrogen
pathway and some controls--from sheepshead minnows. The genes had been previously
identified by differential display reverse transcriptase-polymerase chain
reaction, a method that screens thousands of RNA messages to identify genes
that are turned on or off by specific treatments. The team used microarray analysis
to discover which of the 30 preselected genes were significantly changed by
exposure of fish to estrogenic compounds. They also measured changes in levels
of gene expression when fish were exposed to different concentrations of 17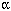 -ethinyl
estradiol, a synthetic estrogen found in birth control pills (which can end
up in waterways via sewer systems).
-ethinyl
estradiol, a synthetic estrogen found in birth control pills (which can end
up in waterways via sewer systems).
Once they had their array in place, the team exposed male sheepshead minnows
to a constant concentration of either strong or weak environmental estrogens.
The strong estrogens included 17ß-estradiol (the normal estrogen found
in vertebrates), 17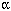 -ethinyl
estradiol, and diethylstilbestrol (a synthetic estrogen formerly used to prevent
miscarriage that caused cancer, reproductive tract abnormalities, and infertility
in the children of women who took it). The weak environmental estrogens included
p-nonylphenol (a breakdown product of alkylphenol ethoxylates, which
are used in various products as washing and cleaning agents, emulsifiers, wetting
agents, and foaming and foam-reducing agents) and the organochlorine pesticides
methoxychlor and endosulfan. Single-stranded DNA for the 30 genes was bound
to multiple membranes.
-ethinyl
estradiol, and diethylstilbestrol (a synthetic estrogen formerly used to prevent
miscarriage that caused cancer, reproductive tract abnormalities, and infertility
in the children of women who took it). The weak environmental estrogens included
p-nonylphenol (a breakdown product of alkylphenol ethoxylates, which
are used in various products as washing and cleaning agents, emulsifiers, wetting
agents, and foaming and foam-reducing agents) and the organochlorine pesticides
methoxychlor and endosulfan. Single-stranded DNA for the 30 genes was bound
to multiple membranes.
To analyze genes that were differentially expressed in the livers of control
and treated fish, the team extracted mRNA and converted it to cDNA, which during
this process was labeled by the addition of a tracer amount of radiolabeled
nucleotides. The cDNA was then incubated with the membranes and bound proportionately
to the 30 genes present thereon. The intensity of the radioactivity in the spots
was directly related to the amount of mRNA present in the sample and, when compared
to controls, was used to determine whether the expression of a gene was elevated
or decreased as a result of exposure to the EDC.
There was an increase in expression of certain genes as a result of exposure.
One endocrine receptor (ER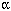 )
was upregulated by every test compound. Four genes involved in the formation
of egg cells were upregulated by every compound except endosulfan. A gene that
plays
)
was upregulated by every test compound. Four genes involved in the formation
of egg cells were upregulated by every compound except endosulfan. A gene that
plays
image credit: PhotoDisc |
| Monitoring mimics. A new model
system profiles the expression of genes affected by exposure to environmental
chemicals--such as those in birth control pills--that may disrupt the endocrine
system. |
an important role in blood clotting also was upregulated by the same five compounds.
Interestingly, the gene for ubiquitin-conjugating enzyme 9, whose metabolic
role is to tag enzymes that have completed their cellular functions and defective
proteins for removal from the cell, was upregulated by p-nonylphenol.
The expression of three genes involved in other processes was downregulated
by the five compounds. Exposure to different concentrations of 17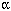 -ethinyl
estradiol also revealed that the microarray method is dose-sensitive, and that
exposure thresholds vary for different genes. These findings could enable calculation
of gene-dependent dose-response curves for evaluating the seriousness of
chemical contamination in environmental cleanup efforts.
-ethinyl
estradiol also revealed that the microarray method is dose-sensitive, and that
exposure thresholds vary for different genes. These findings could enable calculation
of gene-dependent dose-response curves for evaluating the seriousness of
chemical contamination in environmental cleanup efforts.
The scientists plan to expand the expression profiling method to compounds
that mimic other reproductive hormones such as androgen and progesterone. They
are also going to make microarrays for different game fish species used for
food as well as other fish species that are used as standards for monitoring
environmental chemicals.
One hurdle for this technology is obtaining reproducible results. Successful
replication depends on the accuracy of the DNA amplification of each gene, the
correct identification of which genes are bound to the membrane and where, and
the RNA extraction efficiency, because RNA degrades rapidly and can become contaminated
with DNA. These technical steps also require careful laboratory techniques and
multiple replicate experiments to ensure consistent results.
As the methodology expands to include more genes, chemicals, and organisms,
the management and analysis of huge volumes of data will become another hurdle.
Bioinformatics will become increasingly important as these EDC expression profiling
data sets expand. This genetic biomarker assay is an exciting application of
genomics tools for toxicology with promise for finding genes that are affected
by EDCs, for understanding mechanisms that lead to disease, for applying that
knowledge to environmental monitoring and cleanup, and for the rational design
of new compounds that will be safer for human health and the environment.
Mary Eubanks
Metals Leave
Their Mark
Fingerprints
of Low-Dose Exposure
As the emerging field of toxicogenomics continues to progress, the search
for biologically relevant biomarkers of exposure, effect, and susceptibility
is in full swing. Much of the current work focuses on the genomic effects of
potentially toxic metals. In this issue, Angeline Andrew and her colleagues
at Dartmouth College report the results of their study of four metals--arsenic,
cadmium, chromium, and nickel--that have been associated with a variety of adverse
health effects [EHP
111:825-837].
They identified "fingerprints" of early changes in gene and protein expression
in response to each metal that may someday serve as biomarkers of exposure to
these toxicants.
The team used cDNA microarrays to compare the effects of each metal on the
expression of 1,200 human genes in human bronchial BEAS-2B cells. These cells
were chosen because inhalation is a common route of exposure for the metals
currently under study.

image credit: Christopher G. Reuther/EHP, Artville
|
| Fingerprint findings.
New research shows signature effects of four metals on gene expression at
low-dose exposures. This information could lead to molecular biomarkers
of metal exposure. |
In order to ensure that the effects seen were not nonspecific responses to
toxic high doses, the cells were exposed to low, relatively nontoxic doses of
the metal compounds sodium arsenite, cadmium chloride, sodium dichromate, and
nickel subsulfide for 4 hours. They also administered a much higher, cytotoxic
dose of sodium arsenite to explore the effects of dose.
Although the results showed that each of the exposures modified expression
of only a small subset of the 1,200 genes, the data suggest that each metal
modifies expression of a largely unique set of genes that may be characteristic
of each substance. This supports the potential for the development of metal-specific
biomarkers.
There was some overlap in which genes were modified, but none were affected
by all five chemical treatments, and only one, heat shock protein 90A, was modified
by four of the five. Conversely, the authors found it remarkable that the genes
that were altered by more than one treatment were all modified in the same direction,
with either increased or decreased expression. They say this lends support to
the idea that these represent biologically relevant responses to these treatments.
Comparison of the effects of the low and high doses of arsenic also yielded
some unexpected insights. Of a total of 158 genes modified, only 16 were altered
at both doses, and substantially more genes were modified by the lower dose
than by the higher one.
The lower dose modified expression of a wide variety of genes representing
a diverse range of protein classes, such as transcription factors, inflammatory
cytokines, kinases, and DNA repair proteins. The higher dose showed what the
authors call a "striking shift" in the profile, modifying a variety of heat
shock proteins and other genes involved in stress response pathways.
The researchers suggest that this dramatic contrast in gene expression profiles
represents a switch from a survival-based biological response at the lower dose
to a cell death-inducing apoptotic response at the higher dose. Whereas
the high dose of arsenic clearly induced a stress response, it was a more universal,
less toxicant-specific response; the lower doses of the four metals produced
"a more subtle modification of cell signaling pathways," implying a unique,
identifiable signature in the gene expression profile generated by each chemical.
The authors conclude that these metal response patterns may shed new light
on the mechanisms of human diseases caused by toxic metal exposures, and may
also be useful for developing molecular biomarkers of exposure and effect in
future mechanistic, epidemiologic, and risk assessment studies.
Ernie Hood
[Table
of Contents]
Last Updated: May 12, 2003





