Expression and Immunity
New Findings on Ahr Interactions
The transcription factor aryl hydrocarbon receptor (Ahr) can activate
a multitude of genes that regulate the growth and division of cells, including
processes leading to cancer. To date, little is known about the biochemical
processes that mediate the activation of Ahr receptors in the body,
or about the endogenous ligands that bind to and activate these receptors.
This month, researchers from the University of Louisville, Texas A&M University,
and the University of New Mexico, under the leadership of Charles D. Johnson,
report on microarray analyses that illuminate the complexity of Ahr interactions
and thus provide a focus for future experiments [EHP 112:403-412].
These results include a previously unknown relationship between Ahr and
genes involved in the activation of the immune system.
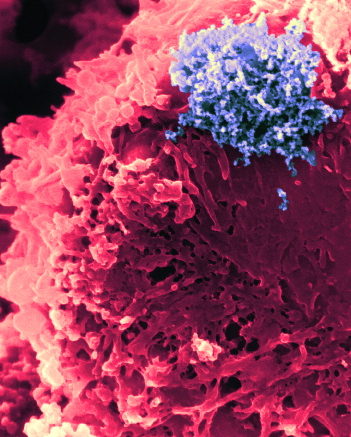 |
T with a twist. Adding an unexpected new element to the map of
relationships between Ahr and retinoids, recent microarray analysis
reveals a novel link between this gene and T cells (above [red], engaging
with a virus [blue]).
image credit: ©Dennis Kunkel Microscopy |
The researchers modeled the identification of relevant components of the biological
response to Ahr ligands using transcriptional profiles of cells from
murine embryonic heart, kidney, and vascular smooth muscle cells. The data
were analyzed using methods developed at the Texas A&M University Genomic
Signal Processing Laboratory to decipher multivariate, nonlinear relationships
among genes.
The analysis focused on 200 of the 12,000 clones on the microarrays that
showed the greatest changes in response to hydrocarbon exposures. The changes
in each of these genes were quantified in relationship to five target genes,
selected because they are known to respond to ligands of Ahr. The first
target gene was Ahr itself. The second was Cyp1b1, which is involved
in the synthesis of steroids.The third was Igfbp-5, which regulates
the functions of the bones, kidneys, and mammary glands. The fourth was Lox,
which transcribes a copper-containing enzyme involved in the creation of connective
tissue. And the fifth was Opn, which transcribes osteopontin, a protein
involved in the mineralization of bone and a component of human atherosclerotic
plaques.
Activity in all five target genes was related to some degree; when one was
expressed, so were the other four. The analysis was not designed to determine
whether activated genes were upstream or downstream of each other--in other
words, which of a given set of genes initiated a reaction.
The expression of Ahr was most closely linked to that of lymphocyte
antigen 6, locus e, which is involved in the activation of T cells. This relationship
had not been predicted by previous studies and helps fill a gap in the map
of relationships among Ahr and retinoids (forms of vitamin A). Lymphocyte
antigen 6e responds to the presence of retinoic acid, and recent research has
shown that Ahr controls the expression of genes that metabolize this
compound. Thus, computational strategies allowed the delineation of connections
between Ahr and retinoic acid that otherwise could not have been predicted
in the absence of biological information.
In other relationships found in this study, Cyp1b1 was best predicted
by the gene that codes for spleen tyrosine kinase, which participates in signaling
leading to activation of the transcription factor NF-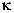 B. Igfbp-5 was
most frequently predicted by Opn; Lox was best predicted by lymphocyte
antigen 6 complex, locus H; and Opn was most often predicted by brain-derived
neurotrophic factor, interleukin 6, and proliferin.
B. Igfbp-5 was
most frequently predicted by Opn; Lox was best predicted by lymphocyte
antigen 6 complex, locus H; and Opn was most often predicted by brain-derived
neurotrophic factor, interleukin 6, and proliferin.
The authors write that the computational approach they used allowed them
to begin constructing gene networks that define broad-ranging interactive biological
relationships. "Although the biological bases for these theoretical relationships
must be investigated further," they conclude, "the number of possible combinations
is now reduced to a manageable size that can be systematically scrutinized
using established molecular methodologies."
Kris Freeman
Taking Stock of Toxicogenomics
Mini-Monograph Offers Overview
Many early concerns about the utility of genomics technologies have largely
been put to rest, but several issues remain to be resolved if toxicogenomics
is to live up to its full potential. Chief among these is the concern that,
although there is a healthy spirit of scientific collaboration and sharing
within the toxicogenomics community, standardized submission of and open access
to data has not yet been accomplished. The International Life Sciences Institute
Health and Environmental Sciences Institute (HESI) Committee on the Application
of Genomics to Mechanism-Based Risk Assessment has been actively engaged in
working on these challenges by sharing experience, best operating practices,
and data to achieve standardization of toxicogenomics data. In this issue,
committee members William Pennie, Syril Pettit, and Peter Lord present an overview
of the committee's research program [EHP 112:417-419]. The overview
by Pennie and colleagues leads into the mini-monograph appearing in this issue,
which explores the issues, challenges, and triumphs of using genomics in mechanism-based
risk assessment.
Established in 1999, the committee is a collaborative research effort incorporating
the unique perspectives and scientific talents provided by its members from
government, academia, and corporate organizations representing the pharmaceutical,
agrochemical, chemical, and consumer products industries. The committee has
conducted and analyzed toxicogenomics experiments within the broad fields of
hepatotoxicity, nephrotoxicity, and genotoxicity with two goals in mind: first,
to determine whether known mechanisms of toxicity could be associated with
characteristic gene expression profiles; and second, to identify technological
and biological sources of variability associated with toxicogenomic experimental
protocols. The answers to these questions helped the committee evaluate the
usefulness of gene expression technology for the purposes of risk assessment.
 |
Working together. Researchers are sharing experience, best operating
practices, and data to achieve standardization in toxicogenomics data.
image credit: Chris Reuther/EHP, Rupesh P. Amin, et al. [EHP 112:465-479
(2004)] |
There is an ongoing need for standardized submission of and open access to
data. Researchers also must have access, through public repository databases,
to standardized microarray data formats that are linkable to toxicology data.
To address these needs, the committee has developed a database in partnership
with the European Bioinformatics Institute. Based on the ArrayExpress database
structure and Minimum Information About a Microarray Experiment (or MIAME)
data format standards, Tox-MIAMExpress will be available to the public early
in 2004, and will contain all of the data generated by the committee's research
collaborations.
According to Pennie and coauthors, the committee's experimental programs
have in fact shown that toxicogenomics is an inherently valuable tool for assessing
toxicity. Specifically, they assert that the value of toxicogenomics is supported
by the research, which has shown four things. First, patterns of gene expression
relating to biological pathways are robust enough to allow insight into the
mechanisms of toxicity. Second, gene expression data can provide strong information
on topographic specificity. Third, dose-dependent changes can be observed.
Finally, concerns about oversensitivity of the technology may be unfounded.
The writers stress that it is important that microarray data results be considered
along with other biologic end points to understand the mechanisms underlying
toxicity. Pathway-level results will be much more relevant for meaningful risk
assessment than single gene expression data, they write, particularly within
the regulatory arena.
Clearly, toxicogenomics has come a long way in the short time since the HESI
committee was first established. As the mini-monograph in this issue shows,
the committee's research portfolio and collaborative approach have contributed
significantly to rapid progress in the field. In all likelihood, this contribution
will continue, thanks to what the authors call "this collective experience
for the benefit of the regulators and regulated industries as well as for the
toxicology community as a whole."
The broad impression imparted by the papers, as expressed by Pennie and colleagues
in their overview of the program, is that "genomics, and more specifically
toxicogenomics, can no longer be regarded as 'new' technology." With increasing
experience has come increasing awareness that toxicogenomics is fast maturing.
The field has proven its value with solid research and significant additions
to the scientific knowledge base; its utility in mechanism-based risk assessment
is less likely to be considered tentative, potential, or pending.
Ernie Hood
[
Table of Contents]
Last Updated: March 9, 2003





