Brief Description
In collaboration with NIA, HPCIO
develops and enhances tools for the archival, retrieval, and mining of genetic
association study data. The Genetic Association Database (GAD) is an
archive of human genetic association studies of complex diseases and disorders.
GAD enables scientists to query association data in a systematic manner and to
integrate association data with other molecular databases. Study data are
recorded in the context of official human gene nomenclature with additional
molecular reference numbers and links. The goal of this project is to collect
all published genetic association study data and allow the user to rapidly
identify medically relevant polymorphism from the large volume of polymorphism
and mutational data, in the context of standardized nomenclature.
PubMatrix SE, is a Web-based text-mining tool on MEDLINE citations. It applies natural language processing and statistical methods on biomedical literature text to provide an estimation of the strength of associations among various entities, including genes and diseases. The results are represented in a matrix format, facilitating more efficient interpretation of large amount of text data to assist in microarray studies.

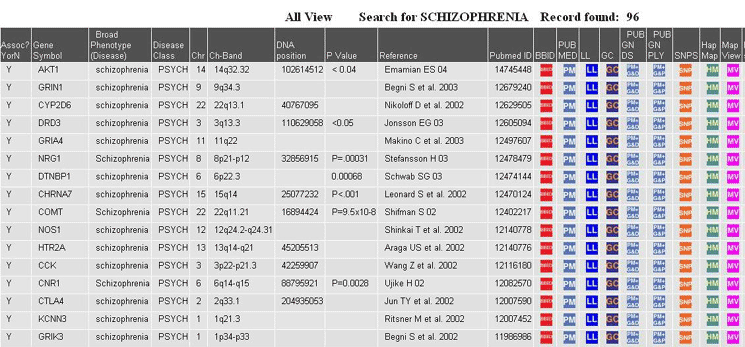
Caption: A simple search of
positive associations for the disease
schizophrenia. Fields in this view include Official Gene Symbol,
Disease
Phenotype, Disease Class, Chromosome, Chromosome Band, Genomic DNA
Position, P
Value, Reference, PubMed ID and Links to other gene related resources

Caption: Search results for
“Candidate Genes for ALZHEIMER
DISEASE” from SNPs3D web site. GAD data have been integrated in the
search results. Each “Y” represents one positive association record
and each “N” represents one negative association record in GAD
Caption:
GAD
links were provided by multiple NCBI applications including Entrez.
User can
access GAD data by following the LinkOut resources for each gene.
List of Collaborators
Major Accomplishments in FY 2007
In collaboration with NIA, HPCIO constantly
improves the quality and quantity of GAD. GAD data has gone through major update
several times in FY2007 to correct missing or erroneous data. The total number
of records has increased from 28 thousands to 40 thousands. In collaboration
with the
A copy of the MEDLINE citations was imported to a local database. The gene
normalization algorithm used in the PubMatrix SE was entered into the
BioCreative II challenge. The competition was an international event to assess
the state-of-the-art text-mining techniques for bioinformatics. Our algorithm
was placed within the top half out of 20 groups. A standalone Web application,
called GIANT, was developed to normalize gene mentions in free-text. This work
is a part of a larger effort, involving a number of institutions in the
Anticipated Major Accomplishments in FY 2008
In FY 2008, HPCIO will create new features to make integrating GAD data more convenient for outside biomedical databases. For example, GAD will add UMLS unique concept identification number (UCI) for each disease. Since current GAD does not store UMLS UCI, some users have to manually associate GAD disease to UMLS UCI before integrating GAD data with their own data. In order to maintain quality and consistency of GAD data, we shell make UMLS UCI available in the future.
Scientific Impact Statement
Albino Bacolla and his colleague from
Metrics
|
40,339 |
|
1,238,655 |
|
49,494 |
|
420 |
|
207,200 76.2% |
Publications in FY 2007
Lau, W.W. and
Lau, W.W. and Johnson C.A. “Rule-based
Human Gene Normalization in Biomedical Text with Confidence Estimation,” in
Proc of Comput Syst Bioinformatics Conf.
2007.
Kevin G Becker, Yonqing Zhang, Narmada Shenoy, Kayla E Smith, Donna Karolchik, Fan Hsu, S Alex Wang, “GAD and GADview: a genomic view of common human disease,” HUGO's 12th Human Genome Meeting, Montreal, Canada, Mon 21-Thu 24 May 2007
Other Publications
Becker, K.G., Barnes, K.C., Bright,
"The Genetic Association Database" Nature Genetics 36:
431-432 (2004)
[Full article pdf] [PubMed]
Sun G, Lau W, Wang A, Shenoy N, Becker K, Cheung H "Ranking and Presenting Gene-Disease Associations from Biomedical Literature." Poster. 2006 Summer Research Program Student Poster Day. [Full Article]
Citations in FY 2007
Han A, Kim WY, Park SM, “SNP2NMD: a database of human single nucleotide polymorphisms causing nonsense-mediated mRNA decay,” Bioinformatics. 2007 Feb 1;23(3):397-9. Epub 2006 Nov 22.
Frodsham AJ, Higgins JP, “Online genetic databases informing human genome epidemiology,” BMC Med Res Methodol. 2007 Jul 4;7:31
Nobuhara Y, Usuku K, Saito M, Izumo S, Arimura K, Bangham CR, Osame M. “Genetic variability in the extracellular matrix protein as a determinant of risk for developing HTLV-I-associated neurological disease,” Immunogenetics. 2006 Jan;57(12):944-52. Epub 2006 Jan 10.
Lussier YA, Liu Y., “Computational approaches to phenotyping: high-throughput phenomics,” Proc Am Thorac Soc. 2007 Jan;4(1):18-25.
Jegga AG, Gowrisankar S, Chen J, Aronow BJ. “PolyDoms: a whole genome database for the identification of non-synonymous coding SNPs with the potential to impact disease,” Nucleic Acids Res.2007 Jan;35(Database issue):D700-6. Epub 2006 Nov 16.
Bacolla A, Collins JR, Gold B, Chuzhanova N, Yi M, Stephens RM, Stefanov S, Olsh A, Jakupciak JP, Dean M, Lempicki RA, Cooper DN, Wells RD, “Long homopurine*homopyrimidine sequences are characteristic of genes expressed in brain and the pseudoautosomal region,” Nucleic Acids Res. 2006 May 19;34(9):2663-75. Print 2006.
Anil G. Jegga, Jing Chen, Sivakumar Gowrisankar, Mrunal A. Deshmukh, RangaChandra Gudivada, Sue Kong, Vivek Kaimal, and Bruce J. Aronow, “GenomeTrafac: a whole genome resource for the detection of transcription factor binding site clusters associated with conventional and microRNA encoding genes conserved between mouse and human gene orthologs,” Nucleic Acids Res. 2007 January; 35(Database issue): D116–D121.
Simon N. Twigger, Mary Shimoyama, Susan Bromberg, Anne E. Kwitek, Howard J. Jacob, and the RGD Team, “The Rat Genome Database, update 2007—Easing the path from disease to data and back again,” Nucleic Acids Res. 2007 January; 35(Database issue): D658–D662.