A Meeting of the Minds on Mice
If genetics research is ever to fulfill its promise of revolutionizing medicine, genotypes must be linked to phenotypes—that is, individual genomic characteristics must be identified and associated with outcomes in the forms of disease susceptibility and/or development; individual responses to drugs, infectious agents, or environmental exposures; or other individual characteristics such as behavioral tendencies. Why does the person who never smoked develop lung cancer, while the three-pack-a-day smoker remains healthy? Why do some people become addicted to drugs, while other users are never hooked? Why does a particular medication work wonders in some people, but not work at all in others? These and countless similar questions represent the enormous challenge still facing researchers as they strive to make personalized medicine a clinical reality.

|
|
Culling out complex traits. A consortium of international scientists is launching new mouse research initiatives to help elucidate genetic components of complex human diseases.
image: PhotoDisc, Arnold Greenwell/EHP, Matt Ray/EHP |
The answers to many of these questions may yet be discovered in the genomes of mice, our diminutive mammalian relatives. That's certainly the hope and belief of the members of the mouse genetics community, 180 of whom gathered 6–9 May 2006 in Chapel Hill, North Carolina, for the fifth annual meeting of the Complex Trait Consortium (CTC), a loosely woven international organization tightly knit in its dedication to elucidating human characteristics by identifying their genetic counterparts in mice.
The "complexity" of complex traits derives from the fact that they are polygenic—multiple genes interact to cause these conditions, and the genes involved may not interact additively. Ninety-three reports presented at the CTC meeting updated progress in the hunt for the multiple genes and quantitative trait loci (or chromosomal "hot spots") associated with a wide variety of complex traits such as heart failure, tumor resistance, obesity, drug and alcohol addiction, and schizophrenia. Sponsored by the NIEHS, the UNC–Chapel Hill, and Agilent Technologies, the conference brought together a diverse group of mouse geneticists, molecular biologists, statisticians, and bioinformaticists from 10 countries.
The CTC is all about collaboration and interaction. "It's unquestionably the best meeting that I go to every year," says Abraham Palmer, an assistant professor of human genetics at the University of Chicago. "The opportunities to communicate with other geneticists working in other fields and with the people who develop our methodology are critically important, and accelerate by months or even years the rate at which the field can move forward," he says.
Karlyne Reilly, a principal investigator in the Mouse Cancer Genetics Program at the National Cancer Institute, agrees. "It brings together a wide variety of science around techniques and how you solve the problems that are common to these different areas," she says. "I always come away with new tools to play with, that I can apply to my own research."
Building a Better Mouse Line
The CTC is presently at the midpoint of building a resource that should prove enormously valuable in the effort to associate genotypes with phenotypes. The Collaborative Cross (CC) is a carefully planned and controlled mouse recombinant inbreeding program that began in 2005 with eight genetically heterogeneous strains. Upon its expected completion in about four years, 1,000 lines closely modeling the breadth of human genetic diversity will have been generated. According to conference keynote speaker Jean-Louis Guénet, a professor emeritus of mouse genetics at the Institut Pasteur in Paris, it will be "one of the most important pages in the book of genetics of the future."
Armed with several powerful new bioinformatic and biostatistical tools being developed specifically to take full advantage of the resource, the CC will enable researchers to hunt far more precisely and efficiently for the multiple genes and quantitative trait loci that constitute complex traits, and will allow the community to share and integrate their raw data sets far more effectively.
"The idea is to accumulate as much diverse data as possible for relatively fixed strains, what we call the 'genetic reference population,'" says Robert Williams, a professor of anatomy and neurobiology at the University of Tennessee Health Science Center and one of the founders of the CTC. "The hope is that everybody will use their own tools—their own methods and their own phenotypes—but the Collaborative Cross will provide a way to bind those results together by using the same animal resource."
According to conference co-organizer David Threadgill, an associate professor of genetics at UNC–Chapel Hill, the goal is for the CC to evolve to become "the central resource for experimental mammalian biology." With a fixed genetic reference population and common tools, he says, "it will be the resource that everybody turns to," because every piece of data collected through the CC will be immediately comparable to any other piece of data in the database.
CC Riding
The CC will enable a so-called systems genetics approach, as opposed to the traditional, laborious effort to identify one gene at a time. As Guénet points out, diseases that are the consequences of the alteration of a single gene—one example is cystic fibrosis—tend to be marginal in terms of frequency. However, polygenic diseases tend to be much more widespread, he says: "Next door to you, you probably have someone with asthma, dermatitis, or autoimmune disease. . . . So we have to work hard to understand the genetic determinants of these complex diseases, and presumably what we are going to learn from the mouse can be transposed to the human being, because we share ninety-eight percent of our genes with the mouse."
Williams shares Guénet's optimism about the tremendous potential of the CC to shed useful light on common human diseases. "You have to understand the function of the gene and its products in a complex milieu, in a mouse or human—not only a mouse or human, but many different mice and many different humans," he says. "We think [the CC] will provide the resource to do that." He adds that the ability to conduct experimental population-based research with the CC should allow much more comprehensive exploration of the genetics associated with gene–environment interactions.
That exploration will also be enhanced by the completion of a mouse genetics initiative undertaken by the NIEHS and Perlegen Sciences to identify the genetic variants in 15 diverse strains of laboratory mice, including SNPs (single-nucleotide polymorphisms), indels (insertions/deletions), and haplotypes (blocks of related SNPs). The database, a project of the recently established NIEHS Center for Rodent Genetics, is scheduled to be unveiled in September 2006, and is anticipated to be a rich and robust source of information for the mouse genetics community.
Signs of early but significant progress in the CC initiative were among the highlights of the meeting. Conference co-organizer John E. French, an NIEHS research physiologist, is encouraged by results emerging from pilot studies. "There's at least been a proof of principle established that it's going to be a very effective tool," he says. "We are only seeing the beginning evidence of that—there's a long way to go—but some of the promise has been identified and, I think, validated." According to Williams, the pilot project is now of sufficient size (two recombinant inbred sets, LXS and BXD, with 80 member strains) that "it provides the community with a good flavor of what this will look like when we have an order of magnitude more strains than we do now."
Threadgill is excited by the flavor that's already emerging. "The major things that are starting to come out are the results of integrating data sets, integrating genetic variations, and integrating gene expression patterns," he says. The new knowledge that's coming out of that—the identity of new genes that are potential master modulators of genetic networks, and how those may actually also be very important for mediating disease processes—speak to the remarkable potential that will be realized when the CC is completed.
A Case in Point
Research results presented by Palmer on his group's work at the University of Chicago illustrate the broad outlines of the types of studies being undertaken by mouse geneticists. Palmer and colleagues are investigating the genetic underpinnings of susceptibility or resistance to drug addiction; given today's working definition of "the environment," recreational drug use is fast becoming a xenobiotic exposure of great interest. An understanding of the genotypic differences between addiction susceptibility and resistance could lead to new targets for therapeutic drugs or preventive interventions.
The team selectively bred mice to have very high or very low sensitivity to locomotor stimulation, a particular behavioral effect of methamphetamine that is a characteristic animal response to drugs of abuse. They then measured the expression of more than 14,000 genes in a region of the animals' brain known to be involved in response to the drug. Ultimately, they arrived at a candidate gene that was found to be very differentially expressed in the high- and low-sensitivity mice—casein kinase 1 epsilon (Csnk1e). It was a gene already known to be involved in locomotor stimulant response of animals to various drugs. But the question then became, was it important in humans?
Fortuitously, thanks to colleague Harriet de Wit of the University of Chicago Department of Psychiatry, Palmer had access to DNA from a cohort of 100 healthy human volunteers. In a double-blind study, the subjects received 0-, 10-, and 20-mg doses of amphetamine in a randomized order. Responses were measured by standardized questionnaires, and were then compared to results of genotyping tests, to see whether there was a correlation between response to the drug and polymorphisms in Csnk1e.
"We found a statistically significant association between this gene, Csnk1e, and people's sensitivity to the euphoric effects of the drug," says Palmer. "So the people with one genotype 'got a buzz,' while people with another genotype didn't. We hypothesize that that may have implications for the likelihood of a person with one genotype who samples the drug to continue to use the drug, and that of course would put them at grave risk for developing an abusive relationship with the drug."
Palmer suspects that polymorphisms in Csnk1e may also be important in a variety of other systems whose mechanisms might be similar to that of addiction. These include the manic phase of bipolar disorder and the use of stimulants to treat attention deficit/hyperactivity disorder.
"I think we're now at a point where it's just about to become easy to go from a phenotype to identifying some of the genes that are involved in that phenotype," says Palmer. "To get all of them is going to take longer, and it's going to require further refinements in our methodology, but I actually think that the story I told is going to become a common story. . . . In the same way that molecular biology took a long time to mature, and now is unbelievably central to the way we think about the progress of medicine and health sciences, I think this field of genetics is right at that turning point."
Knowledge gained from the genomes of our mammalian cousins by groups like the CTC may provide the vital information to eventually usher in the much-anticipated era of personalized medicine. Says Threadgill, "What it really comes down to is being able to predict which individuals are going to be susceptible to certain environmental exposures or disease processes, which individuals are going to respond adversely to combinations of alleles, so that interventions and preventive medicine can be applied where they need to be applied, rather than in global fashion."
Ernie Hood
Environmental Polymorphism Registry
Banking DNA to Discover the Source of Susceptibility
Walking outside on a day when ozone levels are at "code orange" doesn't bother one person, but for someone else, it can result in chest pain, coughing, wheezing, or lung and nasal congestion. Why? Polymorphisms, tiny interindividual variations in genes, may be part of the reason. Providing a pool of information to help researchers determine how these variations interact with the environment to cause disease is the ultimate goal of the Environmental Polymorphism Registry (EPR), which is sponsored by the NIEHS and conducted in collaboration with the University of North Carolina at Chapel Hill's General Clinical Research Center.

|
|
A little prick for a big cause. The Environmental Polymorphism Registry aims to collect 20,000 blood samples that will be studied to help determine how genetic differences may result in disease.
image: Dona McNeill/NIEHS |
Any North Carolinian over 18 years of age can donate a sample—about a tablespoon of blood—to the EPR. Rather than recruiting donors with a particular disease, the EPR aims to gather, over five years, samples from 20,000 people who represent the general state population. The regional nature of the effort facilitates recruitment and follow-up.
"Recruitment is monitored to ensure that the EPR population is representative of the North Carolina population," says Patricia Chulada, one of the four principal investigators of the EPR and a health scientist administrator at the NIEHS. "If we see deficiencies in certain groups, then we can increase efforts targeted to those particular groups."
This approach will help researchers find out which polymorphisms are most common. "We want to look at people's genetic material and find variations, and then go back and figure out what those variations mean," says another EPR principal investigator, Paul B. Watkins, a professor of medicine at UNC–Chapel Hill and director of the General Clinical Research Center.
Chulada and Perry Blackshear, the NIEHS director of clinical research, initiated the registry by approaching Watkins and Susan Pusek, director of training and career development at the General Clinical Research Center. Watkins says the institute—and Blackshear himself—realized that "this is an essential direction of research to understand why some people are healthy and some are sick."
There are multiple DNA registry efforts in the United States. Two major DNA banking efforts include Northwestern University's NUgene Project and the Marshfield Clinic's Personalized Medicine Research Project, both launched in 2002. International DNA banks are even more common, Chulada says. For instance, Iceland's deCODE Project has recruited more than 80,000 subjects and has published findings on genes associated with arthritis and many other common conditions.
The EPR is unique, however, because it is designed to focus on environmentally responsive genes—those that increase the risk of disease when combined with an environmental exposure. The registry was created with the express intent of facilitating clinical studies of polymorphisms in these genes. Being affiliated with the NIEHS, where scientists are already studying such interactions, makes the EPR a natural resource for these investigations.
Protecting Participants
Unlike with anonymous DNA databases, EPR donors provide their names and contact information so they can be asked to participate in follow-up studies if their DNA contains a polymorphism of interest. Participation in follow-up studies is optional, and donors can drop out of the database at any time.
Donors learn about the steps taken to ensure confidentiality in a 6-page consent form. Study interviewers at recruitment tables also discuss this information with potential donors, Chulada says.
Donors' names and other information are stored separately from samples. When a sample is collected, it's assigned a personal identification number. The code key that links the sample to identifying information is kept separate from the sample and from all other data in a computer system that's password-protected. Access to this system is limited to only a few people directly involved in the EPR. Researchers can obtain contact information for potential participants only after approval by the EPR Oversight Committee.
To receive samples, researchers must sign a material transfer agreement, in which the researcher's institution agrees to several conditions. "They can only use the samples for what they outlined in the agreement," Chulada says. "They can't give the samples to others. And they have to destroy the samples within a certain amount of time [which varies on a case-by-case basis]."
In addition, the NIH has granted the EPR a Certificate of Confidentiality, which protects researchers from being required, even by subpoena, to disclose research data or other information about an individual to an outside party such as an insurance company, an employer, or a civil or criminal court. "This is another layer of protection built into this system," Chulada says.
Stepping Up Recruitment
The EPR has already accumulated about 4,000 samples—not far behind the 5,000 collected by NUgene since its launch. The EPR's goal of 20,000 samples is the minimum needed to conduct certain types of studies with adequate statistical power, Chulada says. For example, if a researcher was interested in a rare genetic variant that occurs in only 1% of the population, the variant should be present in 200 samples from a registry of 20,000. "That would give us adequate statistical power to test for a phenotypic association of low to moderate effect, depending on other factors," Chulada says.
When the EPR began, it recruited exclusively at two clinics at UNC–Chapel Hill. It has since expanded recruitment to Rex Hospital in Raleigh and is applying for approval to recruit at Duke University Medical Center in Durham. However, Chulada says, "Although recruiting at medical clinics gave us a diverse population in terms of health and other characteristics, we learned that we could increase both recruitment rates and diversity by recruiting outside of the clinic setting."
A recruitment fair held for five days at the NIEHS campus in Research Triangle Park yielded about 420 donors. "We were ecstatic with the response of the NIEHS community," Chulada says. The general public also can donate through study drives at corporations and health fairs in Research Triangle Park. Potential participants can visit the EPR website (http://dir.niehs.nih.gov/direpr/) to find out about upcoming drives.
A DNA Goldmine
John Hollingsworth, a scientist working in the Environmental Lung Disease Group in the NIEHS Laboratory of Respiratory Biology, is one of the first investigators to apply for use of EPR samples. Hollingsworth and colleagues want to identify people who have a polymorphism in a certain gene, Toll-like receptor 4 (TLR4), known to be important in innate immune responses.
TLR4 was first identified as a candidate gene for response to ozone by NIEHS scientist Steven Kleeberger, leader of the Environmental Genetics Group in the Laboratory of Respiratory Biology. Subsequently, Hollingsworth and colleagues have demonstrated that mice deficient in TLR4 are protected against airway hyperresponsiveness after exposure to ozone. "We want to determine if this gene is important in people in the biologic response to inhaled ozone," Hollingsworth says. "We're trying to validate what we've seen in mice in a human cohort."
Hollingsworth calls the EPR a "goldmine." He says, "It's a perfect situation. We have a cohort willing to be genotyped, rather than doing a mass screening of people for a single project, which is what we've had to do in the past."
Angela Spivey
Beyond the Bench
Continuing Education for Nurses on Environmental Genetics and Complex Diseases
Many people find it hard to fit professional development and continuing education into their busy work lives. Now help is just a mouse-click away for nurses seeking flexible, self-paced training in the growing field of environmental genetics. The Community Outreach and Education Core (COEC) of the Center for Environmental Genetics at the University of Cincinnati, in collaboration with the Genetics Education Program for Nurses (GEPN) of Cincinnati Children's Hospital Medical Center, has created an online Environmental Genetics and Complex Diseases educational module that introduces nurses to the principles of environmental genetics, and also teaches them how to apply those principles in nursing practice.

|
New age of nursing. An online continuing education module introduces nurses to principles of environmental genetics and shows them how to put such principles into practice.
image: geotrac/iStockphoto |
Online since December 2005, the module is useful for all nurses in clinical practice, but especially targets those who work extensively with minority or medically underserved patients. The module focuses on alcoholism, lead, and asthma, three challenging public and environmental health problems in underserved communities.
"The module is designed to prepare nurses in underserved communities to identify people who are at risk for environmental genetic conditions and help those people gain access to community services that emphasize prevention and early treatment strategies," says COEC director M. Kathryn Brown.
Cynthia Prows, a clinical nurse specialist in genetics and the principal investigator of the web program, says the module organizes information into useful and manageable resources. "There is a tremendous amount of information on the Internet about genetics and about environmental health. But how do nurses who have limited knowledge in the topic areas locate the various sites, sift through all the information, decide what information is current and accurate, and then use that information for learning purposes? The answer is, most nurses don't because they don't have the time or the necessary foundational knowledge in genetics to mine the overwhelming mass of information that is accessible through the Internet."
The module developers have done that work for the nurses, and have organized the content in a way that helps nurses develop foundational knowledge in environmental genetics using high-quality resources that are applicable to their practice. Once learners create a unique username and password, they can access the module free of charge, and can re-enter it at any time at the place they last exited. Those who wish to earn 4.8 nursing continuing education contact hours after completing the module and associated evaluations pay a minimal processing fee.
The module offers nurses background information on gene–environment interactions, and teaches them environmental and sociodemographic risk factors for common diseases. It also provides screening tools and community resources for nurses treating patients with recognizable genetic and environmental risk factors. Each of the three learning tracks also offer prenatal, pediatric, and adult case studies and self-assessments with each content area.
After completing the module, nurses are able to approach their communities armed with valuable knowledge of gene–environment interactions and insight into how those interactions can affect human health. They are also equipped with a wealth of online resources that can be accessed long after they complete the training module.
"Making sense of the fast-growing literature about how the health impacts of environmental exposures through the life span are mediated by our genetics is a challenge for health care professionals," says Brown. "We hope that the vast array of resources identified in these self-paced, online modules will be helpful to primary care practitioners trying to make sense of new developments in genetic screening tests, environmental prevention strategies, and treatment options."
The module is available at http://gepn.cchmc.org/. Three additional genetics education modules currently available include Promoting Informed Decision-Making about Genetic Testing, Ethical and Social Issues Related to Genetic Testing, and Interpreting Family History. Two new modules are also in the pilot testing phase: Genetics Is Relevant Now––Nurse Views and Patient Stories, and Nurses' Role in Pharmacogenetics/Pharmacogenomics.
Tanya Tillett
Headliners: Autism
NIEHS-Supported Research
 image: Michael Monahan/ShutterStock |
Misfolded Protein Presents Potential Molecular Explanation for Autism Spectrum Disorders
De Jaco A, Comoletti D, Kovarik Z, Gaietta G, Radic´ Z, Lockridge O, et al. 2006. A mutation linked with autism reveals a common mechanism of endoplasmic reticulum retention for the 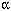 ,β-hydrolase fold protein family. J Biol Chem 281: 9667–9676. ,β-hydrolase fold protein family. J Biol Chem 281: 9667–9676.
Currently, there is only very limited information available on the etiology and biological basis of the autism spectrum disorders, although mutations in the neuroligin 3 and neuroligin 4 genes have caught researchers' attention in recent studies. Now NIEHS grantees Palmer Taylor and Mark H. Ellisman at the University of California, San Diego, and their colleagues have determined, among other findings, that homologous mutations in the genes encoding butyrylcholinesterase (BChE) and acetylcholinesterase (AChE) cause defects in protein processing and expression similar to those seen with neuroligin 3, shedding further light on a potential molecular mechanism underlying autism.
The neuroligins, BChE, and AChE are members of the 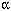 ,β-hydrolase fold family of proteins. The arginine-to-cysteine substitution in the neuroligin 3 mutation was identified in a set of twins, and cell culture studies show most of the expressed protein being retained within the endoplasmic reticulum, suggesting misfolding of the protein. In addition, the small amount of protein that does reach the surface of the cell shows compromised binding affinity for its partner, β-neurexin. Misfolded proteins causing endoplasmic reticulum retention and compromised functional activity are a common consequence of mutations in cystic fibrosis and metabolic diseases with a genetic origin. ,β-hydrolase fold family of proteins. The arginine-to-cysteine substitution in the neuroligin 3 mutation was identified in a set of twins, and cell culture studies show most of the expressed protein being retained within the endoplasmic reticulum, suggesting misfolding of the protein. In addition, the small amount of protein that does reach the surface of the cell shows compromised binding affinity for its partner, β-neurexin. Misfolded proteins causing endoplasmic reticulum retention and compromised functional activity are a common consequence of mutations in cystic fibrosis and metabolic diseases with a genetic origin.
In the current study, confocal fluorescence microscopy and analysis of oligosaccharide processing were used to ascertain whether the homologous arginine-to-cysteine mutation affected AChE and BChE despite their differing oligomerization states. By inserting homologous mutations in the AChE and BChE cDNAs, the investigators found that the mutation also resulted in endoplasmic reticulum retention of the two cholinesterases and enhanced degradation in the proteasome. The authors speculate that altering intracellular oxidation/reduction parameters may influence proper folding and export of these proteins. Jerry Phelps |





