Email comments/bugs to: walthall@stanford.edu
Important: The official site to obtain MW Calc is at PalmGear.com
Be sure to delete the old version before you install the new one.
You now need to install MathLib. Click
here
to get it.
Contents:
Someone made the excellent request that I support Amino Acids.
So, after dragging my feet for far too long, I finally got my act together
last weekend and implemented it. (It turned out to be surprisingly
easy, so I felt really dumb for not doing it sooner.)
What's new in v5.0!
I've had several requests for a new feature since I released this program:
isotope patterns. I finally got around to implementing it!
It unfortunately requires Mathlib since I needed to use pow(...) function
[ie a^b]. You will need to install MathLib. You can get it
here.
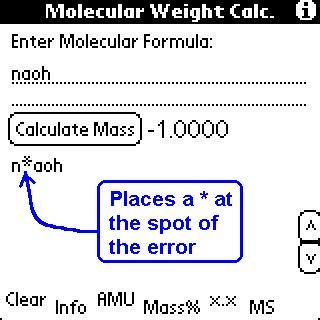


"Grammar" Rules for this program:
(There are certain minor variations, such as Hgs will return the same as HgS, but Hgss will return an error. This is a result of the order of checking that is done on the string. I tried to be as relaxed as possible when checking for error while still ensuring that a correctly inputted formula would return the correct value.)
Correct Usage:
NaOH
HHeLiBe
No
NO
(In the case of No, it would return the mass of Nobelium,
and in the case of NO, it would return the mass of Nitrogen + Oxygen.)
Curly Braces:
{ }
Multiple amino acids can be combined into one set of curly braces. Since both 1 letter codes {AWVG} and 3 letter codes {AlaTrpValGly} are supported, correct capitalization is required for amino acids. In addition, to make it simple to include atoms inside the amino acid braces, parenthesis can be used.
Example:
{AlaGly(CH3)} corresponds to Alanine + Glycine + carbon +
3hydrogens.
When amino acids masses are handled, they are assumed to have had the
HOHremoved.
Thus, {Gly} corresponds to glycine missing an H from
one end of the chain and an OH from the other.
 |
 |
|
|
glycine without the H and OH |
So the string {AlaGly} corresponds in this program to:
To get amino acid sequences that are terminated as normal, one can use many different but equivalent ways:
{AlaGly(H2O)}
{(H)AlaGly(OH)}
{AlaGly}H2O
I prefer the first or second because it reads easier, and it is simpler to later change the nuber of the string. For example, we may want to know what the mass of five alanine-glycine amino acid sequences is. Using the first two above requires only adding the number. The third one above, while faster to write initially, requires either multiplying each part separately by 5, or requires combining them into one set of parenthesis and then multiplying by 5. This is much more prone to error.
{AlaGly(H2O)}5
{(H)AlaGly(OH)}5
{AlaGly}5(H2O)5
({AlaGly}H2O)5
Correct Usage:
na.o.h
h.he.li.br
na....o..h...
.h.f.c.c.o.
na o h
h he li br
Incorrect Usage, but will give the correct answer:
nao.bis.ca
nao bis ca
I would advise that you use the following instead:
na.o.bi.s.ca
na o bi s ca
Incorrect Usage, resulting in an error:
na.oli
na oli
(This would try to find the element Ol, which does not exist.)
Periods and spaces can be used inside of parenthesis as well as inside of curly braces (amino acids).
Correct Usage:
h2.s.o4
H2
H2SO4
H2O
Cl2
.cl2
Incorrect Usage:
h.2.o
2h
PARENTHESIS AND NESTED PARENTHESIS:
[Note that a set of parenthesis can be empty. That is, the following is allowed: Na()OH ]
Correct Usage:
Al(OH)2
.al.(o.h)2
al (o h)2
You can also lookup elements by their atomic number.
Correct Usage using "..":
..al
..h
Incorrect Usage using "..":
..h2co3
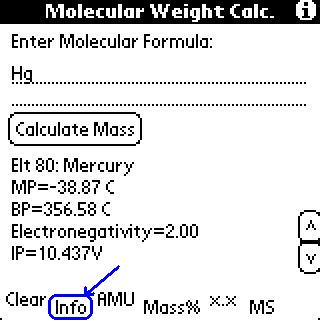
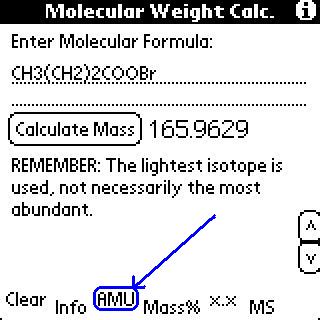
To use the exact masses, click the "AMUs" button.
This program used the most abundant isotope in its calculation
of mass. This feature used the lighest isotope in its calculation
of mass. (This is a change from older versions which did not have
include mass spec stuff.)
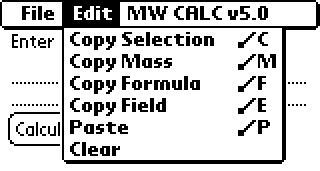
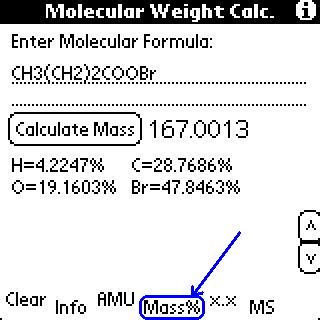
To get mass percents, click the "Mass%" button.
You are limited to ten elements by space constraints of the program.
However, you can copy the extra field "Command:E" or "Copy Field" and paste
it into the notepad to see them all, or use the scroll buttons on the side.
(You are acutally limited to about 30 elements by the length of the string,
but I don't forsee this as a problem. If it is, contact me and I
can make it larger. For people that don't have a program that increases
the size of thier clipboard, they will be limited by their clipboard anyway.)
FUTURE RELEASES:
The users of this program determine what I will implement. If there
are requests for a feature, I'll try to add it. Right now
I am fairly happy with the program, so nothing specific is planned as of
yet.
Here are some features that I plan to implement at some time:
VERSION HISTORY: (b denotes
a beta release, not widely distributed.)
| Version: | Notes about release: |
| 6.0 | Added Amino Acids!
Fixed a display bug: on nested parenthesis, it did not correctly display the spot of the error |
| 5.0 | Added Mass Spec. isotope patterns.
Fixed a minor rounding bug. |
| 4.6 | Added single precision calculations (the x.x
button).
You can now look-up elements by their number or their symbol. |
| 4.0 | Added support for fractional coefficients [ie YBaCuO6.7 ].
Added Mass Percents! Added exact masses (AMUs) for all elements that I could find. ---->Data from Scientific Instrument Services, Inc. Added MP and BP and much other data for elements. ---->Many thanks to PrEpsilon and Craig Counterman's page for the data. |
| 3.0 | Fixed one major bug (the copying of mass/pasting => hard reset bug)
and one minor annoyance (field looks like it has the focus, but doesn't). |
| 2.5 | Added support for spaces. |
| 2.4 | Added support for looking up elements (the CH3Br graphic).
Fixed the ..D and ..T bug. Added the element name when looking up elements (as per user suggestion). (Available to registered users only.) |
| 2.3 | (First widely available release.) Added element lookup. |
| 2.2b | Added nested parenthesis.
Fixed an error where it wrote directly to low memory. |
| 2.1b | Added periods and numbers.
Fixed problem with display. Added Warning and Errors. |
| 2.0.1b | Added the masses of D and T. |
| 2.0b | Ported for PalmIII. Initial beta release. |
| 1.0 | PC version developed. Support for Caps only. |