|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
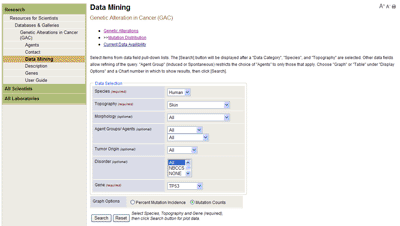
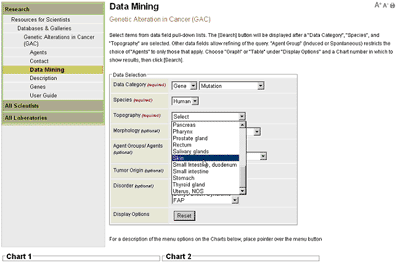
Genetic Alterations in Cancer (GAC)The data are queried using the Data Mining feature described below and the results from each query are displayed in one of four interactive chart areas for comparative analyses. The mutation incidence (percentage of tumors with mutations) from multiple studies of the genes retrieved by the query is displayed in a histogram or listed in a table (user's option). Tables of detailed data from each study, as well as mutation spectra, reference lists for cited studies (with links to PubMed abstracts), and access to study descriptions and related notes are also provided. Data MiningData to be displayed are selected from the pull-down lists provided for each field in the Data Selection window (Figure 1).
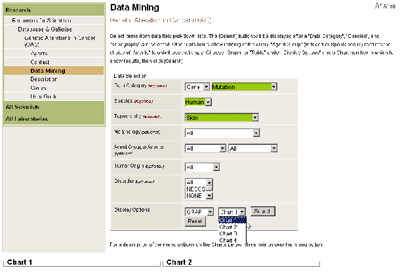
Within Data Mining analysis type, a selection is required in each of the first three fields Data Category, Species, and Topography before a display option and chart area can be selected (see Figure 2). The remaining fields Morphology, Agent, Tumor Origin, and Disorder (human studies) or Strain (rodent studies) are used to refine the query. As selections are made in each data field, options in subsequent fields are restricted to those for which data are available. Display OptionsData can be displayed graphically or in summary tables in any one of the four chart areas. Choose the GRAPH or TABLE display option and a Chart number in which to display query results (Figure 2). Click the [Search] button to start the query. Each chart is independent, and the results are retained until the CLEAR feature on the chart tool bar is selected or a new query is defined for that chart area. Select the Data Mining option on the main tool bar to clear and reset all four charts.
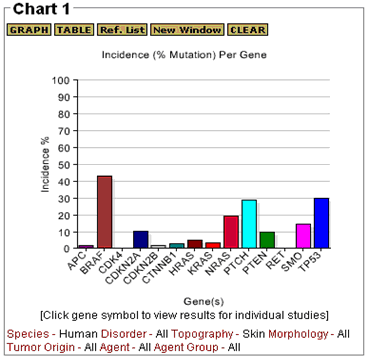
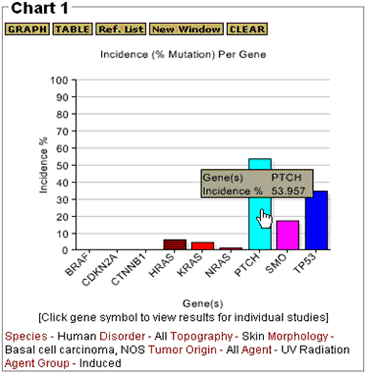
The GRAPH feature generates a histogram that shows the incidence of mutation for each gene that has been evaluated. The TABLE option generates a tabular summary (see Table 1 below) that shows the total number of tumors that have been evaluated, the number of tumors in which mutations were reported, and the incidence (percentage) of tumors with mutations for each gene. Values are calculated based on data from all of the studies represented in the users query. The sample profile plotted in Figure 3 is from a general query for gene mutations in any type of human skin tumor. The results are displayed in Chart 1 and the query selections are shown below the graph. The data include point mutations, frameshifts, and inframe insertions and deletions.
The second sample profile, shown in Figure 4, is from a refined query in which UV Radiation was selected in the agent data field, Basal cell carcinomas for the morphology, and "All" for tumor origin. The profile shows data for all basal cell carcinomas in human skin associated with exposure to UV-radiation. Point to any bar to see the gene symbol and percentage of tumors with mutations.
The TABLE option on the Chart tool bar generates a summary table like that below in Table 1 of the data represented in the graph. The TABLE and GRAPH options are used to toggle between the two views.
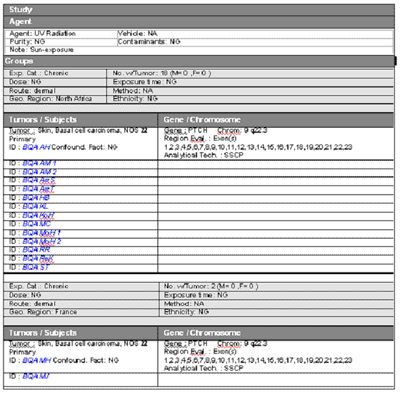
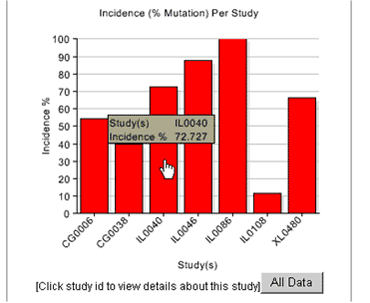
Click any gene symbol in a summary table or bar in a graphic profile to view results for individual studies represented in the display. For example, clicking the bar in Figure 4 for the PTCH gene generates the graph shown in Figure 5. Each study is identified by a Reference ID number. Toggle to the data table (Table 2 below) using the TABLE option on the chart tool bar to see the number of tumors evaluated and the number of tumors that had PTCH mutations in each study. A summary table for each study are also generated by clicking on a gene symbol in Table 1.
A sample data table is shown in Table 2 for the PTCH data. Clicking the CGAP symbol in Table 1 next to any gene displays gene information from the Cancer Genome Anatomy Project (CGAP (http://cgap.nci.nih.gov/)) web site a new window for human or mouse data genes. The symbol RGD beside rat gene symbols links to information on the Rat Genome Database (RGD (http://rgd.mcw.edu/)
Select a study number in the Data Table or a bar in the profile graph to generate a detailed data list for that study. Select the [All Data] button to display data lists for all of the studies shown in the Data Table. A sample data list is shown in Table 3 for the PTCH gene mutation data reported for basal cell carcinomas in reference CG0006. Each entry includes a unique Subject ID number, based on subject identities reported in the reference when available; the subjects gender and age; a field for identifying any known genetic disorder (e.g., xeroderma pigmentosum); an alteration description (Alt. Desc.); the Exon and Codon number for the altered sequence; the codon wild type sequence (WT Seq) and altered sequence (Alt Seq); and the wild type amino acid (WT AA) and altered amino acid (Alt AA) predicted from the altered sequence.
Table 3. Sample Detailed Data List for Results from Study CG0006.
Note that a subject may have more than one mutation reported for a given tumor (see ZPL BCC 05) . If mutations are reported in two or more tumors from the same subject a suffix a, b, c, etc. is added to the Subject ID. Data lists for studies in mice or rats show the strain that was used in place of the Disorder field. The More Detail option on the Data Table tool bar opens a new window and displays detailed study information, including additional agent information, dose and treatment times, identity of the region of the gene that was evaluated (exon numbers), an indication of the analytical technique, and notes for clarification or additional details if needed. A sample display of More Detail is shown in Figure 6 for study number CG0006.
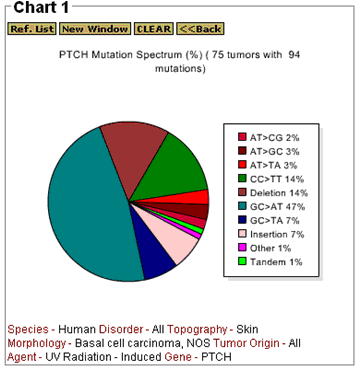
The Gene Mutation Spectrum option on the Chart tool bar generates a pie chart (mutation spectrum) that shows the percentage of each mutation as a portion of the total number of mutations. A sample mutation spectrum is shown in Figure 7 for the PTCH data. The number of tumors with mutations and the total number of mutations reported are shown in the mutation spectrum heading.
The New Window option on the Chart tool bar opens a new, independent, interactive window containing the results displayed in the current chart. Selecting CLEAR on the original chart bar will clear the chart area without clearing the new window. The Ref List option generates a complete bibliographic list of references represented in the active display chart. A sample reference list is shown in Table 4 for the PTCH gene data. Each Reference ID is a unique number that is assigned to a study to identify the data source. The PubMed number, when available, is hyperlinked to its corresponding PubMed abstract.
Table 4. Sample Reference List. s
Mutation DistributionCodon distribution analyses are viewed by using the pull down list for Analysis Type and selecting [Mutation Distribution]. These displays show how base pair substitutions are distributed by codon along the length of different genes. Data Selection OptionsData to be displayed are selected from the pull down lists. For codon distribution analysis, the only fields required before the graph and table areas can be displayed are species and gene highlighted in Figure 8. Other fields can be selected as needed to further refine the query. All selections will remain until either changed or all fields are Reset. As selections are made in each data field, options in subsequent fields are restricted to those for which data is available.
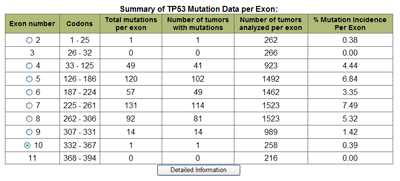
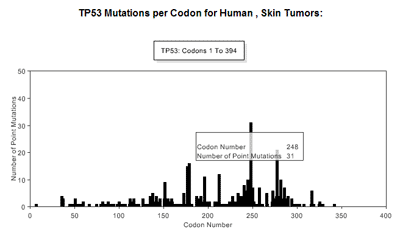
Display OptionsTable/GraphData can be displayed in graphical and/or tabular formats. These are shown when the Display button is selected. The table displays the number of single base substitutions that occur in each exon (Table 5). The codons numbers defining each exon are given by Codon Range, the number of tumors with mutations by Tumors Mutated, and the numbers of tumors analyzed for each exon are also given. The incidence (percent) of mutations that occur in analyzed tumors is also given as % Mutated Tumor Incidence. In addition to this is the option to display the results graphically; this is achieved by selecting the Include at the Display Options, Graph prompt. This feature generates a bar chart that shows the number of single base substitutions that occur at each codon across the length of a gene (Figure 9). Point to any bar to see the codon number and number of base substitutions occurring. Large genes may be divided into separate but contiguous graphs. Values are calculated based on point mutation data from all studies represented by users selections, but excludes tandem and complex substitutions.
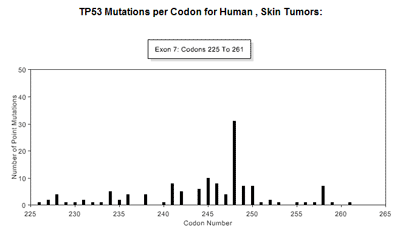
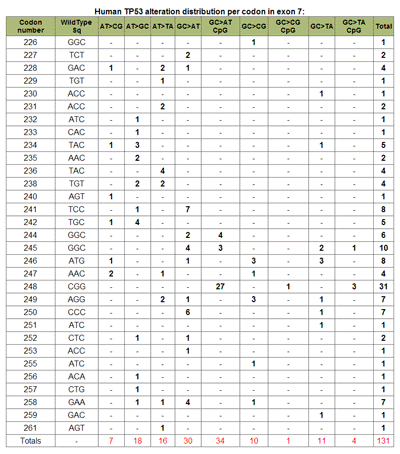
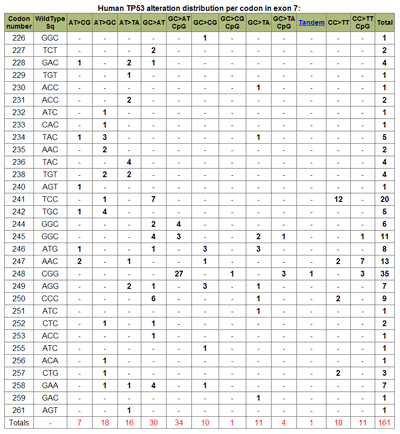
Graph Display TypeThe type of graphic display can be modified by choosing to either display the number of mutations per codon (default), or the incidence of mutations at each codon per exon. This is achieved by choosing the appropriate selection under Display Options-> Graph Display Type. (See Figure 8). Summary TableA summary table describing, in detail, the results presented graphically can be selected at the Display Options -> Summary Table option. This summarizes the codon numbers at which mutations occurred and the number of mutations that were present at each. Detailed InformationFrom the table feature more Detailed Information (see Table 5) about individual exons can be selected using the radio buttons for an individual exon. An expanded graphical display of the distribution of mutations for the selected exon and the types of base pair substitutions occurring is generated. A sample profile of the mutations that occur in Human, Skin, TP53, exon 7 is shown in Figure 10 and Table 6. The bar chart in Figure 10 displays the number of single base substitutions that occur at each codon; Point to any bar to see the codon number and number of base substitutions present.
Tandem MutationsThe displayed data can be further modified to include tandem mutation information with the Data Selection options (Figure 11).
When Include Tandem Mutation(s) and Display are selected both the graph and the table are modified to include all tandem point mutations. In the table these are listed separately as CC>TT, CC>TT CpG, and Tandem. The latter describes any two consecutive base substitutions occurring in 1 codon that are not CC>TT changes including those occurring at CpG sites. The number of non-CC>TT tandem mutations occurring at CpG sites is given as a footnote (Table 7).
The types of substitutions that occur as tandem mutations can be shown by selecting Include Tandem Mutation(s) Table and Display. This produces a table (Table 8) listing the tandem base substitutions that occur by codon number. |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||



 ) web site for the corresponding gene.
) web site for the corresponding gene.