|
CW Barnes1, LJ Szabo1, 2, JL Johnson1, VC Bowersox3, SV Krupa2, DA Gay3, KS Harlin3
1USDA-ARS, St. Paul, MN, 2University of Minnesota, St. Paul, MN
3National Atmospheric Deposition Program, Illinois State Water Survey, Champaign IL.
INTRODUCTION
Atmospheric transport and deposition constitutes the major mechanism for dispersal of certain rust fungi. Trap nurseries, currently used to follow seasonal movements of rusts, are labor intensive and reflective of past events. An early warning system for the arrival of spores is needed. Puccinia graminis was used as a model rust system to test whether real-time PCR can detect rust fungal spores in rain samples. The data were used to create back trajectory models to determine the source of inoculum.
OBJECTIVES
- To develop a protocol for removal of spores from filters.
- To estimate the detection limit of the real-time PCR assay.
- To test filtered rain samples along the Puccinia pathway for stem rust fungal spores through the wheat growing season.
- To use back trajectory models to determine the source region of inoculum.
METHODS
1. Collect spores
|

NADP Collector |
 |
2. Remove spores from filters

Fig. A. Puccinia graminis spores were spotted onto a filter, and visualized at 200X (left). The filter was sonicated for 1 minute at 60 oC (right).
3. DNA extraction. The OmniPrep™ DNA extraction kit was used.
4. Nested real-time PCR assay

Fig. B. Schematic of the real-time PCR assay. Arrows represent primers and the bar represents the TaqMan probe. Primers used were a general fungal primers (F), rust specific primers (R). The TaqMan probe (Pg) is specific for P. graminis.
5. Detection limits
1. The detection limit was determined by spiking filters with a known quantity of spores, with the detection limit of approximately 10 spores per half filter.
2. Dog hair was used to pick single P. graminis spores and placed directly into centrifuge tubes (no filter). One spore was detected 80% of the time.
Distribution of Wheat Along the Puccinia Pathway
RESULTS
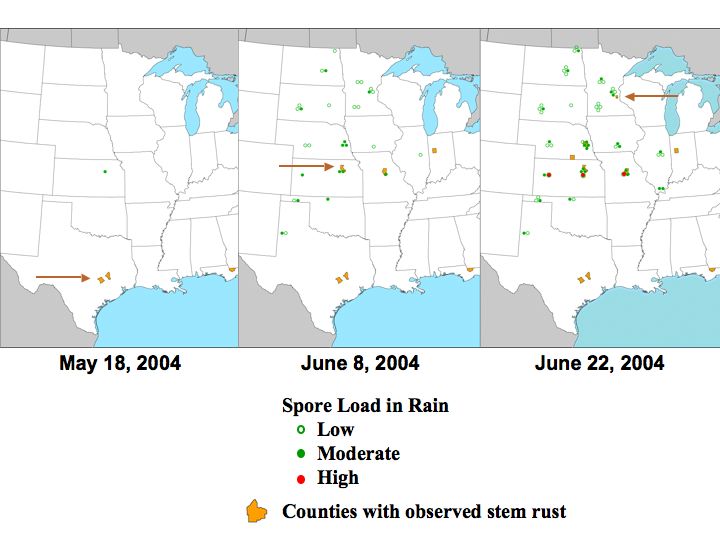
Maps of stem rust progression and spore detection.

Back Trajectory Model

SUMMARY
- Heat and sonication were used to remove spores from rain filters.
- Detection limit of the assay was approximately 10 spores per half filter.
- Filtered rain samples from along the Puccinia pathway all tested positive for stem rust fungal spores.
· Low levels of spores were detected 4 weeks prior to 1st observed infections in the field.
· Moderate levels of spores were detected 3 weeks prior to 1st observed infections in the field.
· High levels of spores were detected by mid June in Kansas and Missouri and may be from local inoculum sources.
- Back trajectory model suggests stem rust spores moved along Puccinia pathway, and traveled relatively long distance.
ACKNOWLEDGEMENTS
Funding was provided by the USDA-ARS CRIS, Minnnesota Soybean Research and Promotion Council, and the Minnesota Agricultural Rapid Response fund. We’d also like to thank Jacki Morrison for graphical assistance.
|