|
VRC
Home
Research Laboratories
Structural Biology Laboratory
Peter Kwong, Ph.D.
 telephone: 301-594-8439
telephone: 301-594-8439
fax: 301-480-2658
e-mail: pdkwong@nih.gov
Address: Vaccine Research Center, NIAID, NIH
40 Convent Drive
Bldg. 40, Room 4508
Bethesda, MD 20892
Group Members (2007):
Priyamvada Acharya, Ph.D.; Lei Chen, Ph.D.; Leo Kong, B.A.;
Young Do Kwon, Ph.D.; Shahzad Majeed, M.S.; Gilad Ofek, Ph.D.;
Marie Pancera, Ph.D.; Anita Shah, Ph.D.; Jonathan Stuckey;
Tongqing Zhou, Ph.D.
Description of Research Program (2007):
The Structural Biology Laboratory seeks to apply structural
biology to the development of an effective HIV-1 vaccine.
Despite the enormous potential of atomic-level design — successfully
used, for example, in the development of the HIV-1 protease
inhibitors — current vaccine development makes little use
of atomic-level information. We are trying to change this.
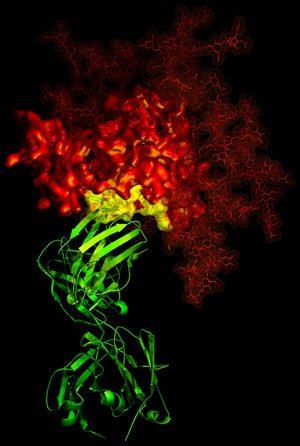
HIV-1 generally manages to evade antibody neutralization.
Its gp120 glycoprotein (red) has one potential weakness,
the site of binding (yellow) for the CD4 receptor. The
b12 antibody (green) exploits this weakness - utilizing
a functional requirement for rapid association with CD4
- to neutralize HIV-1. This image, from X-ray crystallography,
catches the b12 antibody in the act of grabbing onto this
site of vulnerability. |
One area in which we and others have already
made an impact is in understanding how HIV-1 is able to evade
the humoral immune system. Determination of the structure
of the HIV-1 gp120 envelope glycoprotein, the primary target
of neutralizing antibodies against HIV-1, showed how N-linked
carbohydrate can form both an immunologically silent face
— with carbohydrate masquerading as "self" — and also can
protect neighboring epitopes through an "evolving glycan shield."
We also showed how conformational flexibility of gp120 can
combine with quaternary restrictions within the viral spike
to prevent antibody neutralization. These and other studies,
defining for example protective variable loops, have led to
an understanding of the molecular trickery that protects HIV-1
from the humoral immune response.
But can one use structural biology in vaccine
design? Currently, we are following two lines of investigation.
One line involves the precise delineation of
functional constraints to identify potential footholds of
conservation and exposure. In a collaborative study, we investigated
antibodies that bound to the co-receptor-binding site on gp120
and found them capable of neutralizing not only HIV-1, but
even the more evolutionary divergent HIV-2. We found such
that CD4i antibodies develop to high titers in most HIV-1
infected individuals. Unfortunately, our analysis also found
that the virus hides the site of co-receptor binding so that
— prior to engagement of the primary HIV-1 receptor, CD4 —
the co-receptor site is not formed.
These studies demonstrate the strength of functional
constraints in restricting epitope variation. But they also
identify an important weakness: functional conservation does
not necessarily engender epitope exposure, which is required
for antibody neutralization. We are currently exploring how
function might constrain the site of CD4 binding, which —
unlike the co-receptor-binding site — must be available as
an initial site of attachment.
A second line of investigation involves structural
analysis of the few broadly neutralizing antibodies that have
been identified and that have the ability to neutralize diverse
isolates of primary HIV-1. Only four antibodies of such ability
have thus far been identified, the antibodies 2F5, 2G12, 4E10,
and b12.
We have determined the structures of both 2F5
and also of b12, each with their HIV-1 envelope epitopes.
In a collaborative study, primarily with David Baker's group
at the University of Washington, Joe Sodroski's group at the
Dana-Farber Cancer Institute, and also with Gary Nabel's and
Rich Wyatt's groups at the Vaccine Research Center, we are
currently generating epitope mimics, unencumbered by known
mechanisms of humoral evasion. Tests of these epitope mimics
in small animals should reveal their potential to elicit antibodies
similar to the template broadly neutralizing ones.
Whether the confluence of structural information
that we are generating is sufficient to elicit broadly neutralizing
antibodies will depend in part on our ability to iteratively
optimize immunogenicity and also on structural parameters
of conformational mimicry, epitope accessibility, elicited
potency, neutralization breadth, and target specificity. Our
investigations have already led to insight into the parameters
governing antibody elicitation and neutralization. True success,
however, will depend on whether or not we succeed in creating
immunogens capable of substantially reducing the incidence
of HIV-1 infection in humans.
Links:
Description of Research Program
(2001)
Selected publications:
- Zhou T, Xu L, Dey B, Hessell AJ, Ryk DV, Xiang SH,
Yang X, Zhang MY, Zwick MB, Arthos J, Burton DR, Dimitrov
DS, Sodroski J, Wyatt R, Nabel GJ, Kwong PD. (2007)
Structural
definition of a conserved neutralization epitope on HIV-1
gp120. Nature 445, 732-737.
- Douek DC, Kwong PD, Nabel GJ. (2006). The
rational design of an AIDS vaccine. Cell 124, 677-81.
- Huang CC, Tang M, Zhang MY, Majeed S, Montabana
E, Stanfield RL, Dimitrov DS, Korber B, Sodroski J, Wilson
IA, Wyatt R, Kwong PD. (2005) Structure
of a V3-containing HIV-1 gp120 core. Science 310, 1025-8.
- Zhou T, Hamer DH, Hendrickson WA, Sattentau QJ,
Kwong PD. (2005) Interfacial
metal and antibody recognition. Proc Natl Acad Sci U
S A 102, 14575-80.
- Pancera M, Lebowitz J, Schon A, Zhu P, Freire
E, Kwong PD, Roux KH, Sodroski J, Wyatt R. (2005)
Soluble
mimetics of human immunodeficiency virus type 1 viral spikes
produced by replacement of the native trimerization domain
with a heterologous trimerization motif: characterization
and ligand binding analysis. Journal of Virology 79,
9954-69.
- Huang C, Stricher F, Martin L, Decker JM, Majeed
S, Barthe P, Hendrickson WA, Robinson J, Roumestand C, Sodroski
J, Wyatt R, Shaw GM, Vita C and Kwong PD. (2005)
Scorpion-toxin
Mimics of CD4 in Complex with Human Immunodeficiency Virus
gp120: Crystal Structures, Molecular Mimicry, and Neutralization
Breadth. Structure 5, 755-68.
- Decker JM, Bibollet-Ruche F, Wei X, Wang S, Levy
DN, Wang W, Delaporte E, Peeters M, Derdeyn CA, Allen S,
Hunter E, Saag MS, Hoxie JA, Hahn BH, Kwong PD, Robinson
JE and Shaw GM. (2005) Antigenic
Conservation and Immunogenicity of the HIV Co-Receptor Binding
Site. J. Exp. Med. 9, 1407-19.
- Kwong PD. (2005) Refolding
the Envelope. Nature, 433, 815-816.
- Ofek G, Tang M, Sambor A, Katinger H, Mascola
JR, Wyatt R and Kwong PD. (2004) Structure
and Mechanistic Analysis of the Anti-HIV-1 Antibody 2F5
in Complex with Its gp41 Epitope. Journal of Virology,
78, 10724-10737.
- Huang CC, Venturi M, Majeed S, Moore MJ, Phogat
S, Zhang MY, Dimitrov DS, Hendrickson WA, Robinson J, Sodroski
J, Wyatt R, Choe H, Farzan M, Kwong PD (2004).
Structural
basis of tyrosine sulfation and VH-gene usage in antibodies
that recognize the HIV type 1 coreceptor-binding site on
gp120. Proc Natl Acad Sci U S A. 101, 2706-11.
- Kwong PD (2004). The
447-52D antibody: hitting HIV-1 where its armor is thickest.
Structure 12, 173-4.
- Burton DR, Desrosiers RC, Doms RW, Koff WC, Kwong
PD, Moore JP, Nabel GJ, Sodroski J, Wilson IA, Wyatt RT
(2004). HIV
vaccine design and the neutralizing antibody problem.
Nat Immunol. 3, 233-6.
- Xiang S-H, Wang L, Abreu M, Huang C, Kwong PD,
Rosenberg E, Robinson JE and Sodroski J. (2003).
Epitope
mapping and characterization of a novel CD4-induced human
monoclonal antibody capable of neutralizing primary HIV-1
strains. Virology 325, 124-134.
- Kock M, Pancera M, Kwong PD, Kolchinsky P, Grundner
C, Wang L, Hendrickson WA, Sodroski J and Wyatt R. (2003).
Structure-based,
Targeted Deglycosylation of HIV-1 gp120 and Effects on Neutralization
Sensitivity and Antibody Recognition. Virology 313,
387-400.
- Labrijn AF, Poignard P, Raja A, Zwick MB, Delgado
K, Franti M, Binley J, Vivona V, Grundner C, Huang CC, Venturi
M, Petropoulos CJ, Wrin T, Dimitrov DS, Robinson J, Kwong
PD, Wyatt RT, Sodroski J, Burton DR. (2003). Access
of Antibody Molecules to the Conserved Coreceptor Binding
Site on Glycoprotein gp120 Is Sterically Restricted on Primary
Human Immunodeficiency Virus Type 1. J Virol. 77, 10557-10565.
- Majeed, S, Ofek G, Belachew A, Huang C, Zhou T
and Kwong PD. (2003). Enhancing
Protein Crystallization through Precipitant Synergy.
Structure 11, 1061-1070.
- Choe H, Li W, Wright PL, Vasilieva N, Venturi
M, Huang C, Grundner C, Zwick MB, Wang L, Rosenberg ES,
Kwong PD, Burton DR, Robinson JE, Sodroski JG and Farzan
M. (2003). Tyronsine
Sulfation of Human Antibodies Contributes to Recognition
of the CCR5 Binding Region of HIV-1 gp120. Cell 114,
161-170.
- Wei X, Decker JM, Wang S, Hui H, Kappes JC, Wu
X, Salazar-Gonzalez JF, Salazar MG, Kilby JM, Saag MS, Komarova
NL, Nowak MA, Hahn BH, Kwong PD, Shaw GM. (2003).
Antibody
neutralization and escape by HIV-1. Nature 422, 307-312.
- Raja A, Venturi M, Kwong P, Sodroski J. (2003).
CD4 Binding Site Antibodies Inhibit Human Immunodeficiency
Virus gp120 Envelope Glycoprotein Interaction with CCR5.
J Virol. 77, 713-718.
- Kwong PD, Doyle ML, Casper DJ, Cicala C, Leavitt
SA, Majeed S, Steenbeke TD, Venturi M, Chaiken I, Fung M,
Katinger H, Parren PW, Robinson J, Van Ryk D, Wang L, Burton
DR, Freire E, Wyatt R, Sodroski J, Hendrickson WA, Arthos
J. (2002). HIV-1
evades antibody-mediated neutralization through conformational
masking of receptor-binding sites. Nature 420, 678-682.
- Xiang SH, Kwong PD, Gupta R, Rizzuto CD, Casper
DJ, Wyatt R, Wang L, Hendrickson WA, Doyle ML, Sodroski
J. (2002). Mutagenic
stabilization and/or disruption of a CD4-bound state reveals
distinct conformations of the human immunodeficiency virus
type 1 gp120 envelope glycoprotein. J Virol. 76, 9888-9899.
- Sanders RW, Venturi M, Schiffner L, Kalyanaraman
R, Katinger H, Lloyd KO, Kwong PD, Moore JP. (2002).
The
mannose-dependent epitope for neutralizing antibody 2G12
on human immunodeficiency virus type 1 glycoprotein gp120.
J Virol. 76, 7293-7305.
- Yang X, Lee J, Mahony EM, Kwong PD, Wyatt R,
Sodroski J. (2002). Highly
stable trimers formed by human immunodeficiency virus type
1 envelope glycoproteins fused with the trimeric motif of
T4 bacteriophage fibritin. J Virol. 76, 4634-4642.
- Kwong, P. D., Wyatt, R., Majeed, S., Robinson,
J., Sweet, R. W., Sodroski, J. & Hendrickson, W. A.
(2000). Structures
of HIV-1 gp120 Envelope Glycoproteins from Laboratory-Adapted
and Primary Isolates. Structure 8, 1329-1339.
- Myszka, D. G., Sweet, R. W., Hensley, P., Brigham-Burke,
M., Kwong, P. D., Hendrickson, W. A., Wyatt, R., Sodroski,
J., Doyle, M. L. (2000). Energetics
of the HIV gp120-CD4 binding reaction. Proc. Natl. Acad.
Sci. USA 97, 9026-9031.
- Yang, X., Florin, L., Farzan, M., Kolchinsky,
P. Kwong, P. D., Sodroski, J. & Wyatt, R. (2000).
Modifications
that Stabilize Human Immunodeficiency Virus Envelope Glycoprotein
Trimers in Solution. J. Virol. 74, 4746-4754.
- Kwong, P. D., Wyatt, R., Sattentau, Q. J., Sodroski,
J. & Hendrickson, W. A. (2000). Oligomeric
Modeling and Electrostatic Analysis of the gp120 Envelope
Glycoprotein of the Human Immunodeficiecy Virus (HIV).
J. Virol. 74, 1961-1972.
- Moulard, M., Lortat-Jacob, H., Mondor, I., Guillaume,
R., Wyatt, R., Sodroski, J., Zhao, L., Olson, W., Kwong,
P. D. & Sattentau, Q. J. (2000). Selective
Polyanion Interactions with Basic Surfaces on Human Immunodeficiency
Virus Type 1 gp120. J. Virol. 74, 1948-1960.
- Kwong, P. D., Wyatt, R., Desjardins, E., Robinson,
J., Culp, J. S., Hellmig, B. D., Sweet, R. W., Sodroski,
J. & Hendrickson, W. A. (1999). Probability
Analysis of Variational Crystallization and Its Application
to gp120, the Exterior Envelope Glycoprotein of Type 1 Human
Immunodeficiency Virus (HIV-1). J. Biol. Chem. 274,
4115-4123.
- Kwong, P.D. & Liu, Y. (1999). Use
of Cryoprotectants in Combination with Immiscible Oils for
Flash-cooling Macromolecular Crystals. J. Applied Crystallography
32, 102-105.
- Zhang, W., Canziani, G., Plugariu, C., Wyatt,
R., Sodroski, J., Sweet, R. W., Kwong, P. D., Hendrickson,
W. A. & Chaiken, I. (1999). Conformational
Changes of gp120 in Epitopes near the CCR5 Binding Site
are Induced by CD4 and a CD4 Miniprotein Mimetic. Biochemistry
38, 9405-9416.
- Binley, J. M., Wyatt, R., Desjardins, E., Kwong,
P. D., Hendrickson, W. A., Moore, J. P. & Sodroski,
J. (1998). Analysis
of the Interaction of Antibodies with a Conserved Enzymatically
Deglycosylated Core of the HIV Type 1 Envelope glycoprotein
120. AIDS Research Human Retroviruses 14, 191-198.
- Rizzuto, C. D., Wyatt, R., Hernandez-Ramos, N.,
Sun, Y., Kwong, P. D., Hendrickson, W. A. & Sodroski,
J. (1998). A
Conserved HIV gp120 Glycoprotein Structure Involved in Chemokine
Receptor Binding. Science 280, 1949-1953.
- Wyatt, R., Kwong, P. D., Desjardins, E., Sweet,
R. W., Robinson, J., Hendrickson, W. A. & Sodroski,
J. (1998). The
Antigenic Structure of the HIV gp120 Envelope Glycoprotein.
Nature 393, 705-711.
- Kwong, P.D., Wyatt, R., Robinson, J., Sweet, R.
W., Sodroski, J. & Hendrickson, W. A. (1998).
Structure
of an HIV gp120 Envelope Glycoprotein in Complex with the
CD4 Receptor and a Neutralizing Human Antibody. Nature
393, 648-659.
|
Dr. Peter D. Kwong
NIH/Vaccine Research Center
40 Convent Drive
Bldg. 40, Room 4508
Bethesda, MD 20892
|
|



